ISSN: 0973-7510
E-ISSN: 2581-690X
The use of microorganisms for the remediation of petroleum hydrocarbons contaminated environments has been shown to be more suitable and convenient alternative to other approaches. In this study, degradation of aliphatic fractions of Bonny light crude oil by five bacteria: Bacillus cereus ATCC 14579, Campylobacter hominis ATCC BAA-381, Dyadobacter koreensis NBRC 101116, Micrococcus luteus Sn1-1 and Pseudomonas aeruginosa aab51a04 isolated from bitumen-contaminated surface water in Agbabu, Ondo State (E04048-491 and N06034-361) was investigated. The residual aliphatic hydrocarbons after degradation were estimated by Gas chromatography/Mass Spectroscopy analysis. The GC-MS analyses revealed that D. koreensis had the best degrading ability; it reduced 57% of the total detectable aliphatic compounds in the crude oil while Campylobacter hominis had the least percentage reduction (28%). P. aeruginosa, B. cereus and Micrococcus luteus had 42%, 38% and 35% percentage reduction respectively. However, crude oil treated with the combination of the five bacteria resulted in 70% reduction of the total detectable aliphatics thus revealing better degradative performance than individual species.
Biodegradation, Aliphatic hydrocarbons, Crude oil, Gas Chromatography.
Crude oil is composed of several compounds, such as aliphatic, aromatic and polyaromatic hydrocarbons (PAH), sulfur, oxygen and nitrogen containing compounds. The processing of crude oil to usable products is usually accompanied with environmental pollution problems (Okerentugba and Ezeronye, 2003).
This has immensely contributed to the pollution of soil and water (Tri, 2002). Petroleum contaminated soil contains various hazardous materials such as aromatic hydrocarbons and polycyclic aromatic hydrocarbons which are potentially toxic, mutagenic and carcinogenic. Several approaches such as physical, chemical and biological techniques have been developed to unravel the problem of petroleum pollution. As a result of the drawbacks associated with the physicochemical approach, biological alter-natives have received more attention over time. (Okoh, 2006).
Bioremediation, an alternative clean up technology is based on the acceleration and enhancement of the natural processes of biodegradation through the selection, concentration, and acclimation of micro-organisms to degrade many toxic hydrocarbons during a short period of time (Islas-Garcia et al., 2015).
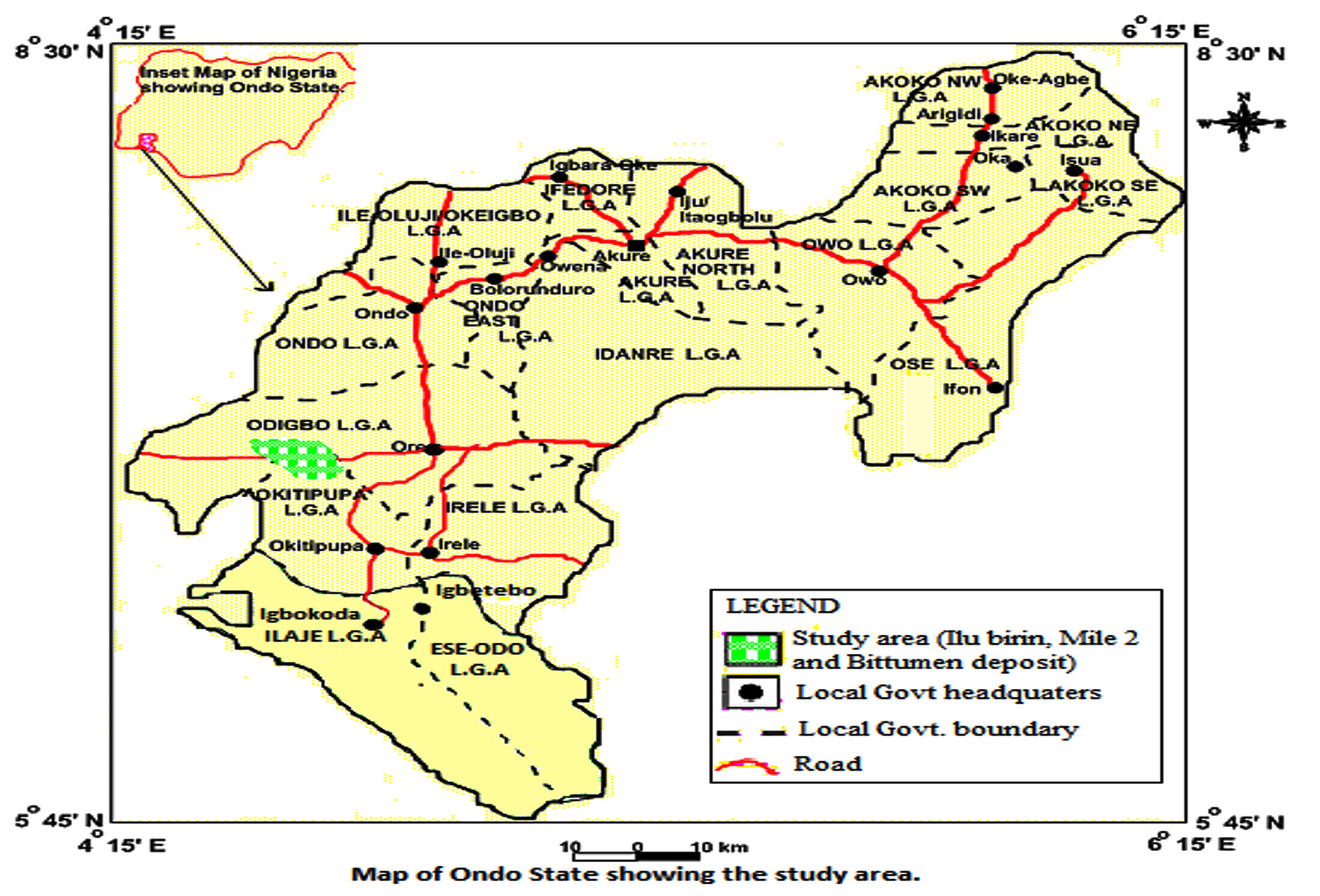
The use of microorganisms for the degradation of petroleum and its products in the environment has been proven as an efficient, cost-effective, flexible and environmentally sound treatment (Margesin and Schinner, 2001). Agbabu is a village of about 8, 611 inhabitants in Ondo State, south-west, Nigeria which has the largest deposit of bitumen and heavy oil. High levels of aliphatic hydrocarbons, polycyclic aromatic hydrocarbons and polychlorinated biphenyls in the bitumen deposit impacted area have been reported by Fagbote and Olanipekun (2010b). Pollution of water with hydrocarbon wastes enhances indigenous microbial populations, which are capable of utilizing the hydrocarbon substrates as their sole carbon and energy sources thereby degrading the contaminants. This study therefore aims at evaluating the degradative capabilities of bacteria indigenous to this bitumen deposit impacted area.
Source of isolates
Five hydrocarbon degrading bacteria isolates from our previous research identified via 16S rRNA sequencing were used for this study. They are Bacillus cereus ATCC 14579, Campylo-bacter hominis ATCC BAA-381, Dyadobacter koreensis NBRC 101116, Micrococcus luteus Sn1-1 and Pseudomonas aeruginosa aab51a04 (Olowomofe et al., 2018).
Degradative Potentials of Single and Mixed Bacteria on Crude oil
The strains previously stored on slant were re-plated on nutrient agar plates before being inoculated into nutrient broth and incubated overnight. The isolates, singly and in consortium were inoculated in mineral salt medium [0.2g of KCl, 6g of Na2HPO4, 2.8g of NaH2PO4, 0.1g of MgS04 and 5g of NaCl] containing 2% v/v crude-oil. The set up was incubated at 30oC for 21days. The optical density (OD 600nm) was measured at seven days intervals using spectrophotometer (JENWAY6705) while the residual crude oil was also extracted at seven days intervals for analysis (Malik and Ahmed, 2012).
Residual Crude oil extraction
Residual oil was extracted by liquid-liquid extraction as described (Minai-Tehrani and Herfatmanesh, (2007). The inoculated samples were mixed vigorously with 50ml hexane: dichloromethane (1:1) in a separating funnel. The mixture was allowed to separate into different fractions, and the oil fraction was carefully collected. The organic layer was separated from the aqueous layer by allowing the funnel with its content to settle on the retort stand rack before being emptied into a 250ml borosilicate beaker.
Total Aliphatic Hydrocarbon separation
The extract was fractionated by liquid-solid chromatography using a column packed with activated silica gel G-60. The column was sequentially eluted with hexane: dichloro-methane (1:1) to yield fractions containing aliphatic hydrocarbons. The mixture was concentrated to 1.0ml by stream of the nitrogen gas before the gas chromatography analysis (Minai-Tehrani and Herfatmanesh, 2007).
Gas chromatography analysis
The aliphatic and aromatic fractions were analyzed as described by the EPA Method 1655 (EPA, 2011). One micro litre (1µL) was injected into Gas Chromatography column through the injection port. Nitrogen was used as a carrier gas and a constant flow rate of 2ml/min was set. Injector and detector temperatures were 250oC and 350oC respectively. The column temperature was programmed at an initial temperature of 70oC; this was held for 2 min, and then ramped at 10oC/min to 320oC and held for 10 min.
Statistical Analysis
Microsoft excel and Chi- square of the Statistical Procedure for Social Science version 22.0 (SPSS, Chicago, IL, USA) were used for the data analysis.
Degradation studies were carried out with five hydrocarbon- degrading bacteria recovered from Agbabu bitumen deposit, Ondo State in our previously reported investigation and the result of the gas chromatography analysis is presented below:
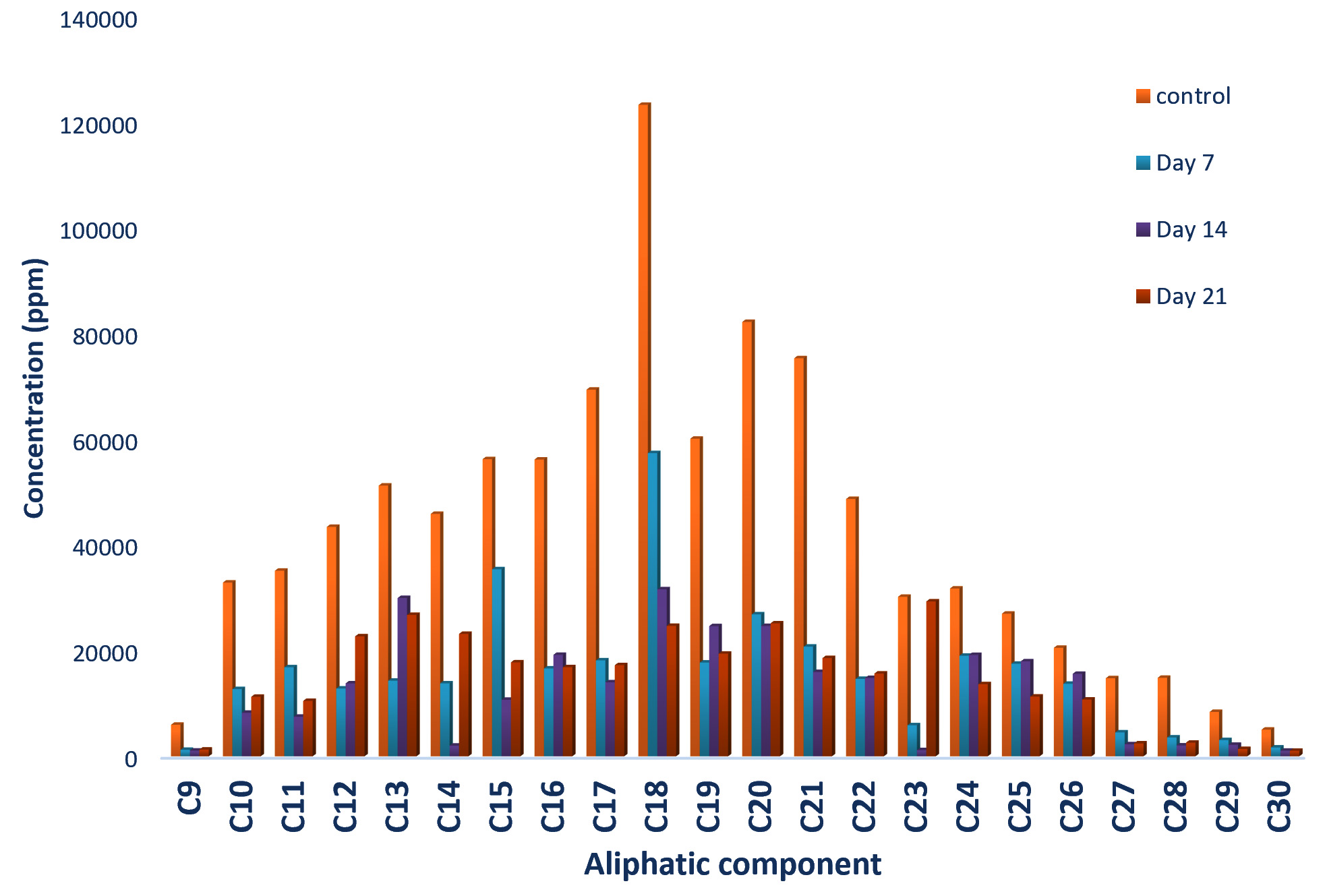
The concentration of aliphatic compounds present in the n-hexane extract of residual crude oil from Bacillus cereus culture fluid is presented in Fig. 2. Twenty-two aliphatic compounds were detected at different concentrations in the control sample; C18 (Octadecane) had the highest concentration of 123215.0 ppm while C30 (Tricontane) had the lowest concentration of 5030.3 ppm. Reduction in the concentrations of the entire aliphatic compound was observed on the 7th day, though statistical analysis revealed that there was no significant difference between their concentrations at control and day 7. As the incubation period increased to 14 day, concentrations of 55% of the aliphatic compounds reduced further while the remaining 45% had increase in their concentrations. Further reduction in their concentrations was observed at day 21 except for C9 Nonane, C11 Undecane, C12 Dodecane, C15 Pentadecane, C18 Octadecane, C21 Henicosane, C22 Docosane, C25 Pentacosane and C28 Octacosane whose concentrations increased.
 Fig. 2.Concentrations of Aliphatic compounds in residual crude oil from Bacillus cereus culture fluid.
Fig. 2.Concentrations of Aliphatic compounds in residual crude oil from Bacillus cereus culture fluid.Pseudomonas aeruginosa was found to degrade most of the aliphatic compounds in the residual crude oil samples with observed reduction of the peaks eluted in the chromatogram as shown in Fig.3. There was reduction in all the aliphatic compounds on the 7th day of incubation while fourteen out of the twenty-two aliphatic compounds detected in the control sample further reduced in concentrations after 14 days but there was noticeable increase by the 21st day. The analysis of variance however revealed that there was no significant difference in their concentrations at control, day 7, 14 and 21.
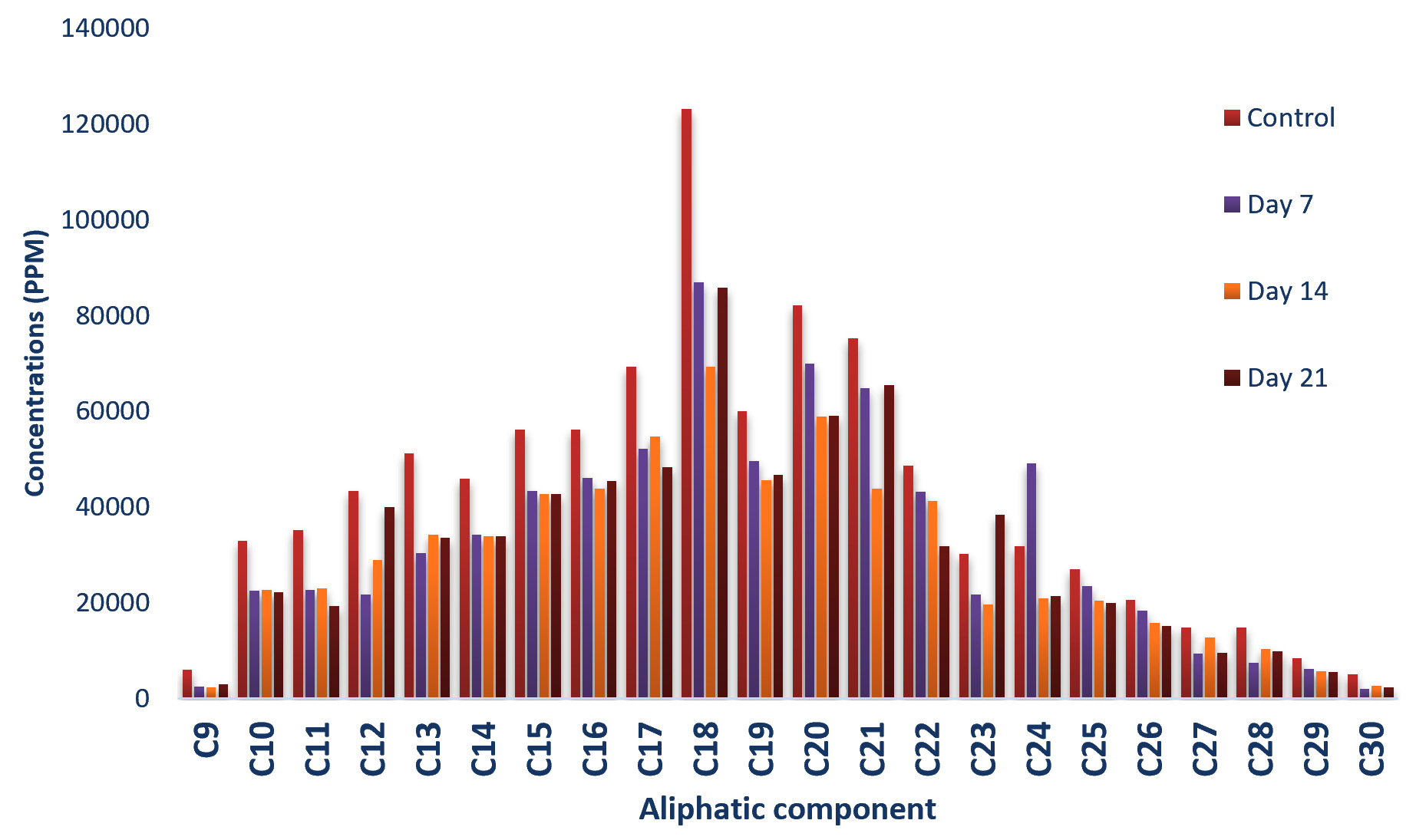 Fig. 3.Concentrations of Aliphatic compounds in residual crude oil from Pseudomonas aeruginosa culture fluid.
Fig. 3.Concentrations of Aliphatic compounds in residual crude oil from Pseudomonas aeruginosa culture fluid.In Fig. 4, the concentration of residual aliphatic components of the degraded crude oil by Dyadobacter koreensis is presented following GC analysis. Statistical analysis showed significant reduction in the concentrations of all the detectable aliphatic compounds after 7 days. Further decrease in their concentrations was observed as concentrations of about 68% of the aliphatic compounds reduced till after 14 days and later increased as the incubation time increased to 21 days while the remaining 32% had increase in their concentrations after the 7th day till 21 days.
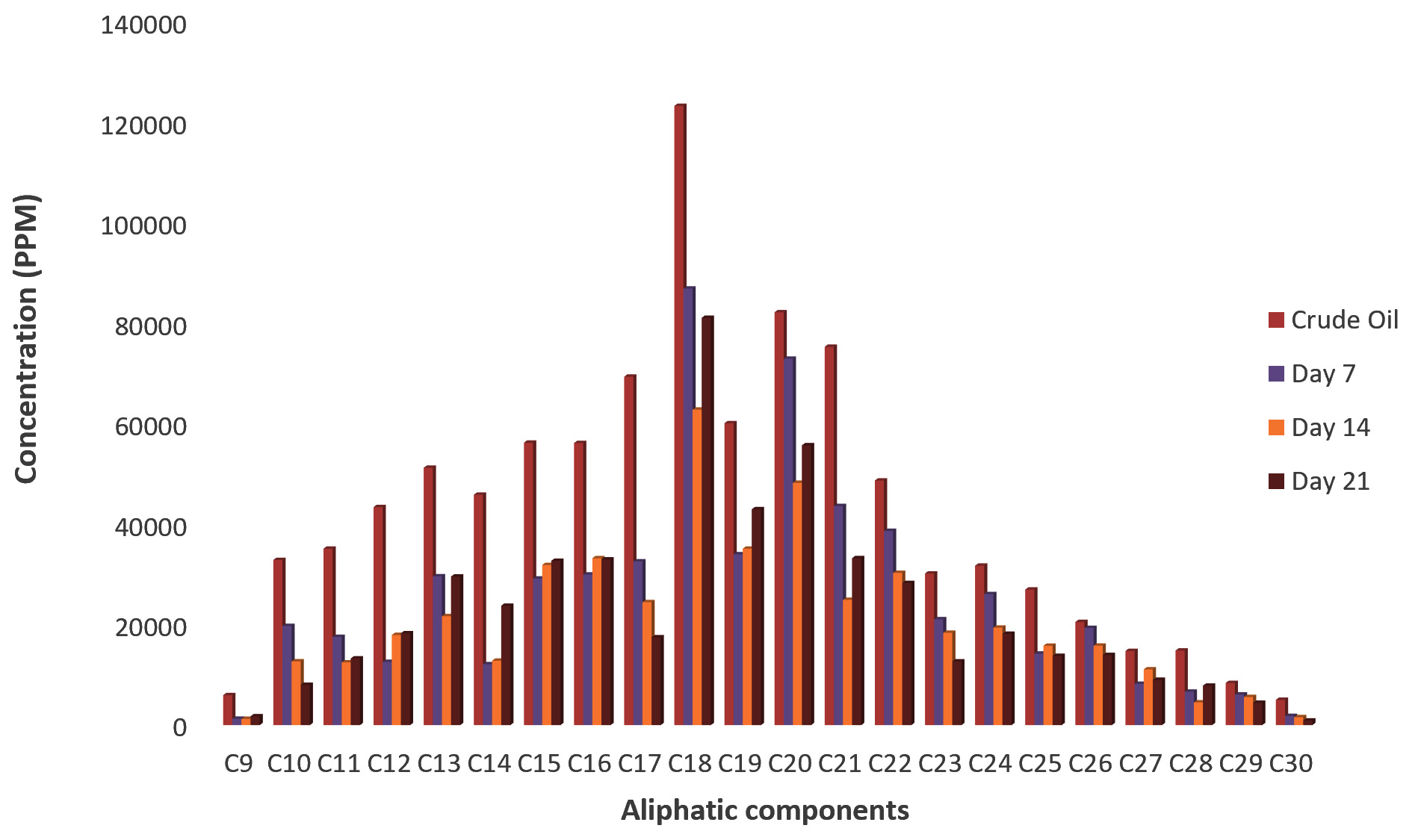 Fig. 4. Concentrations of Aliphatic compounds in residual crude oil from Dyadobacter koreensis culture fluid.
Fig. 4. Concentrations of Aliphatic compounds in residual crude oil from Dyadobacter koreensis culture fluid.Fig. 5 shows the concentrations of aliphatic compounds in residual crude oil from Campylobacter hominis culture fluid. Concentrations of all the aliphatic hydrocarbons also reduced significantly by the 7th day of incubation with a further decrease observed till day 14 except for C11 (Undecane)C14 (Tetradecane), C15 (Pentadecane), C26 (Hexacosane) and C27 (Heptacosane) which increased. However, most of the compounds had their concentrations reduced till day 21 except C9 (Nonane) C14 (Tetradecane), C15 (Pentadecane), C18 (Octadecane), C23 (Tricosane) and C25 Pentacosane where increase in concentration was reported.
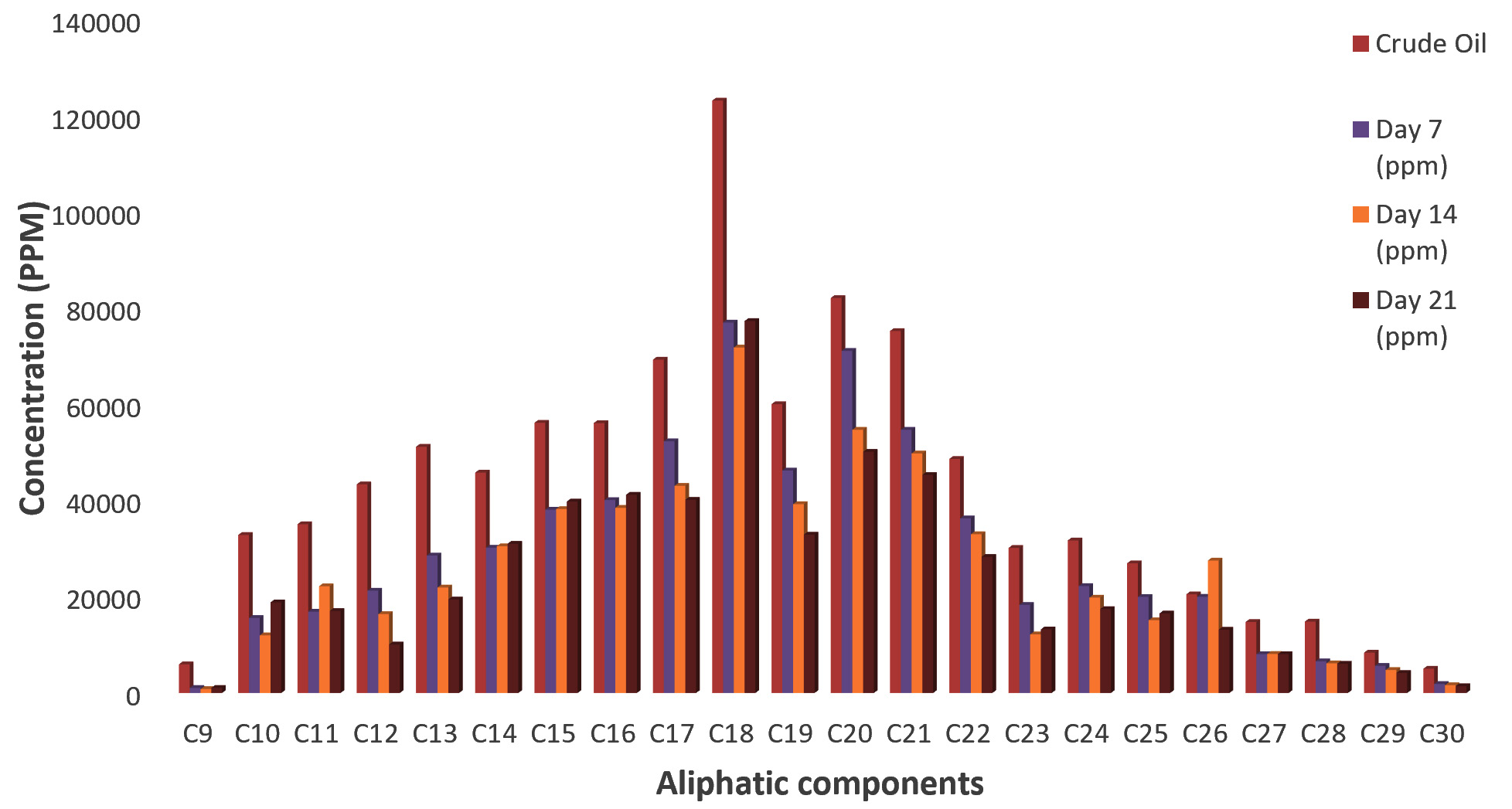 Fig. 5.Concentrations of aliphatic compounds in residual crude oil from Campylobacter hominis culture fluid
Fig. 5.Concentrations of aliphatic compounds in residual crude oil from Campylobacter hominis culture fluidFig. 6 shows the concentration of aliphatic compounds in residual crude oil from Micrococcus luteus culture fluid. The highest reduction in the concentrations of all the aliphatic hydrocarbons was observed on the 7th day of incubation. As the incubation period increased, their concentrations increased. Further reduction in their concentrations was observed on day 14 except C10 (Decane) C11 (Undecane) C12 (Dodecane) C13 (Tridecane) C14 (Tetradecane) C15 (Pentadecane) C22 (Docosane) C23 (Tricosane) C24 (Tetracosane) C26 (Hexacosane) and C28 (Octacosane). Likewise, on day 21, concentrations of aliphatic compounds reduced significantly as compared with that of the control except C9 (Nonane) C16 (Hexadecane) C18 (Octadecane) C19 (Nonadecane) C20 (Eicosane) and C21(Henicosane) which had increased concentrations.
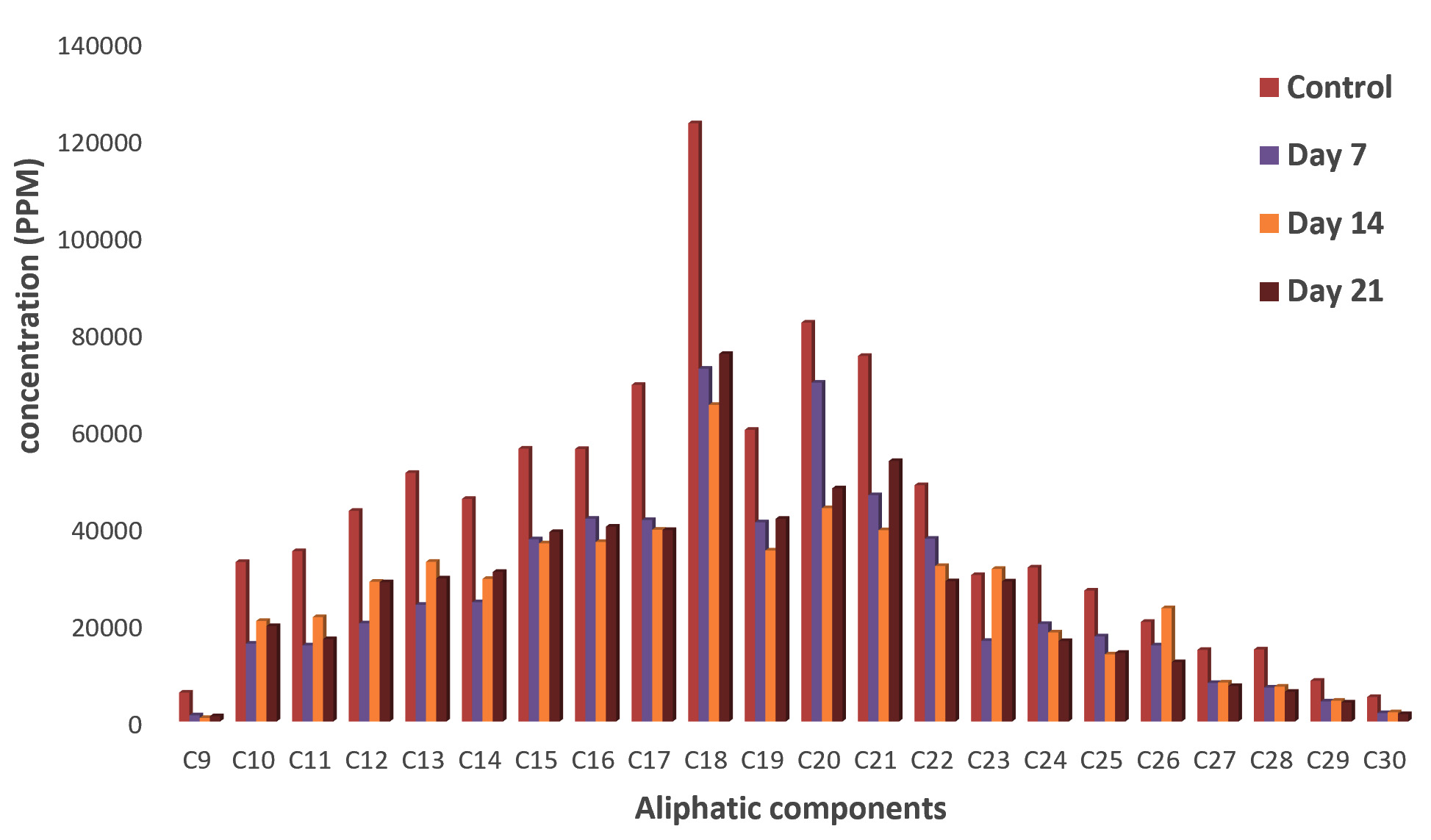 Fig. 6. Concentrations of Aliphatic compounds in residual crude oil from Micrococcus luteus culture fluid
Fig. 6. Concentrations of Aliphatic compounds in residual crude oil from Micrococcus luteus culture fluidInoculation of consortium led to significant reduction in the concentrations of aliphatic compounds in the crude oil. Concentrations of all the aliphatic compounds were significantly reduced on the 7th day of incubation. Further reduction of these compounds was observed on day 14 except C12 (Dodecane) ,C13 (Tridecane) , C14 (Tetradecane) , C16 (Hexadecane) , C19 (Nonadecane) , C22 (Docosane), C24 (Tetracosane), C25 (Pentacosane) and C26 (Hexacosane) which had increased concentration. On day 21, concentrations of C13 (Tridecane), C16 (Hexadecane), C18 (Octadecane), C19 (Nonadecane), C24 (Tetracosane), C25 (Pentacosane), C26 (Hexacosane), C29 (Nonacosane) and C30 (Tricontane) reduced while concentrations of C9 (Nonane) C10 (Decane), C11 (Undecane), C12 (Dodecane C14 (Tetradecane) C15 (Pentadecane) C17 (Heptadecane) C20 (Eicosane) C21 (Henicosane)C22 (Docosane) C23 (Tricosane) C27 (Heptacosane) C28 (Octacosane) increased (Fig. 7).
Petroleum and petro-chemical products are known as major source of environmental pollution in the recent decades (EPA, 2013). Hazardous impact of these pollutants in terrestrial and aquatic habitats cannot be overemphasized. However, the use of auto-chthonous microbes indigenous to hydrocarbon-polluted niches for bioremediation of these pollutants has been broadly recognized as an efficient technique to combat this challenge. (Brooijmans et al., 2009; Gopalakrishnan and Divya, 2014; Esedafe et al., 2015). The organisms are known for their ability to remove these toxic compounds from the environment through production of inducible enzymes, mutations and acquisition of hereditary materials from closely related populations within the hydrocarbon-challenged community (Joshi and Pandey, 2011; Olanipekun et al., 2012). This study evaluated the degradative activities of some bacteria from bitumen polluted surface water.
The bacteria were screened down to five isolates based on their abilities to maintain steady growth in crude oil supplemented medium, as determined by their optical densities and growth rates. The GC/MS analyses revealed different concentrations of aliphatic components from C9 to C30. The major constituent of crude oil is the aliphatic fraction which contains linear, branched and cyclic carbons (Mishra et al., 2001; Bhattacharya et al., 2003; Jain et al., 2005). As indicated in the GC fingerprints of B. cereus treated sample, visible reduction in the peaks and concentrations of these compounds shows the organism was able to mineralize the aliphatic fractions of the crude oil. The pattern of degradation showed that this organism degraded the lower and higher hydrocarbon chains better than those of the middle carbons. This agreed with previous findings that microorganisms metabolized the lower and higher hydrocarbon chains before the middle chains during hydrocarbon degradation (Bello, 2007; Malik and Ahmed, 2012; Al-Sulami et al., 2014). Reduction in concentrations of all aliphatic compounds after 7 days indicated that the organism was utilizing these compounds for its cell metabolism. However, increase in concentrations of some of the compounds on day 14 and day 21 could have resulted from higher aliphatic compounds degraded to lower compounds, thereby increasing the concentration of such lower hydrocarbon chains.
Pseudomonas aeruginosa followed the same degradation pattern as B. cereus but higher reduction in the concentration of the aliphatic components of the crude oil was observed. Previous findings have emphasized the prevalence of Pseudomonas species in hydro-carbon contaminated environments and reported their catabolic competence and genetic agility. Pseudomonas is known to have great affinity for linear and cyclic hydrocarbons. (Vankateswaran et al., 1995; Nojiri et al., 1999; Obayori et al., 2008). Significant reduction in the concentrations of aliphatic compounds in D. koreensis treated sample showed that the organism possessed the ability to use the aliphatic components of the crude oil for growth and metabolism. This organism appeared the best out of the three hydrocarbon degrading bacteria studied due to its ability to reduce significant amount (57%) of the aliphatic compounds compared to the other two organisms. Although the genus Dyadobacter have been isolated from hydrocarbon polluted environment, no study has established its crude oil degrading ability. The catabolic versatility of this organism could be attributed to long term exposure to the pollutants, which resulted in stimulation of suitable degradative genes.
Bacterial Consortium which combined the five hydrocarbon degrading bacteria reduced 70% of all the aliphatic compounds in the crude oil after 14 days. The degradative potential of the consortium was much higher than when the organisms were inoculated singly. There has been perception that a single organism could not metabolize wide range of hydrocarbon substrates (Adebusoye et al., 2007). This has led to the affirmation that microbial conglomerate displayed better degradative proficiency than pure culture strains (Leahy & Colwell, 1990; Adebusoye et al., 2007). Tao et al. (2017) also reported that a defined co-culture of an indigenous bacterial consortium and exogenous Bacillus subtilis effectively speed up the degradation of crude oil.
It has also been reported that degradation of complex hydrocarbons, such as crude oil requires diverse microbial populations with extensive enzymatic capacities. Such mixed cultures display metabolic flexibility and dominance compared to pure cultures (Leahy and Colwell, 1990; Hamme et al., 2000; Juckpech et al., 2012). Thus, a microbial consortium which contains a mixture of microbes that produce appropriate enzymes for degradative pathways is considered to be well suitable for the degradation of hydrocarbons.
Aknowledgements
None
Conflicts Of Interest
The authors declare that there is no conflict of interest.
Authors’ Contribution
All authors have made substantial, direct and intellectual contribution to the work and approved it for publication.
Funding
None.
Data Availability
All datasets generated or analyzed during this study are included in the manuscript.
Ethics Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
- Abass A.A., Abass O.A., Adejare A.A. and Ojeyemi M.D. Distribution of Polycyclic Aromatic Hydro-carbons in surface soils and water from the vicinity of Agbabu bitumen field of south-western Nigeria. Journal of Environmental Science and Health, 2007; 42(8): 1043-1049.
- Adebusoye S.A., Ilori M.O., Amund O.O., Teniola O.D. and Olatope S.O. Microbial degradation of petroleum hydrocarbons in a polluted tropical stream, World Journal of Microbiology and Biotechnology, 2007; 23: 1149-1159.
- Al-Sulami A. A., Al-Taee A.M.R., and Al-Kanany F.N. A. Improving oil biodegradability of aliphatic crude oil fraction by bacteria from oil polluted water. African Journal of Biotechnology, 2014; 13(11): 1243-1249.
- Bello Y.M. Biodegradation of Lagoma crude oil using pig dung. African Journal of Biotechnology, 2007; 6: 2821-2825.
- Bhattacharya D., Sarma P.M., Krishnan S., Mishra S. and Lal B. Evaluation of Genetic Diversity among Pseudomonas citronellolisStrains Isolated from Oily Sludge-Contaminated Sites. Applied and Environmental Microbiology, 2003; 69(3): 1435–1441.
- Brooijmans R.J.W., Pastink M.I. and Siezen R.J. Hydrocarbon-degrading bacteria: the oil-spill clean-up crew. Microbial Biotechnology, 2009; 2: 587-594.
- Cabello M.N. Hydrocarbon pollution: its effect on native Arbuscular Mycorrhizal Fungi (AMF). FEMS Microbiology Ecology, 1997; 22: 233-236.
- E.P.A. Analytical Methods for the Oil and Gas Extraction Point Source Catergory. United States Environmental Protection Agency (US EPA), Engineering and Analysis Division Office of Water, Washington DC., United States. 2011; pp 58-60.
- E.P.A. A citizen’s guide to bioremediation. United States Environmental Protection Agency (US EPA), Washington DC., United States. 2013; pp 38-40.
- Esedafe W.K., Fagade O.E., Umaru F.F. and Akinwotu O. Bacterial Degradation of the Polycyclic Aromatic Hydrocarbon (PAH)-Fraction of Refinery Effluent. International Journal of Environmental Bioremediation and Biodegra-dation, 2015; 3(1): 23-27.
- Fagbote E.O. and Olanipekun E.O. Levels of polycyclic aromatic hydrocarbons and poly-chlorinated biphenyls in sediment of bitumen deposit impacted area. International Journal of Environmental Science and Technology, 2010; 7(3): 561-570.
- Gopalakrishnan R. and Divya C.V. The pattern of degradation of diesel by Acinetobacter species from diesel oil contaminated soil. Scrutiny International Research Journal of Microbiology and Bio-Technology (SIRJ-MBT), 2014; 1(5): 16-23.
- Hamme J.D., Odumeru J.A. and Ward O.P. Community dynamics of a mixed-bacterial culture growing on petroleum hydrocarbons in batch culture. Canadian Journal of Microbiology, 2003; 46: 441-450.
- Hesham A., Khan S., Tao Y., Li D. and Zhang Y. Biodegradation of high molecular weight PAHs using isolated yeast mixtures: application of metagenomic methods for community structure analyses. Environmental Science Pollution Research International, 2012; 19: 3568-3578.
- Holt J.G., Kreieg N.R., Sneath P.H., Staley J.T. and Williams S.T. Bergey’s Manual of Determinative Bacteriology, William and Wilkins Co., Baltimore, 1994; 559-564.
- Jain R.K., Kapur M., Labana S., Lal B., Sarma P.M., Bhattacharya D. and Thakur I.S. Microbial diversity: application of microorganisms for the biodegradation of xenobiotics. Curriculum Science, 2005; 89: 101–112.
- Joshi R.A. and Pandey G. B. Screening of petroleum degrading bacteria from cow dung. Research Journal of Agricultural Sciences, 2011; 2: 69-71.
- Juckpech K., Onruthai P. and Panan R. Degradation of polycyclic aromatic hydrocarbons by newly isolated Curvularia sp. F18, Lentinus sp. S5, and Phanerochaetesp. T20. Science Asia, 2012; 38: 147-156.
- Leahy J.G. and Colwell R.P. Microbial degradation of hydrocarbons in the environment. Microbiology Review, 1990; 54: 305-540.
- Malik Z.A., and Ahmed S. Degradation of petroleum hydrocarbons by oil field isolated bacterial consortium”, African Journal of Biotechnology, 2012; 11(3): 650-658.
- Margesin R. and Schinner F. Bioremediation (natural attenuation and biostimulation) of diesel-oil-contaminated soil in Alpine glacier sking area. Applied Environmental Microbiology, 2001; 67: 3127-3133.
- Megharaj M., Ramakrishnan B., Venkateswarlu K., Sethunathan N. and Naidu R. Bioremediation approaches for organic pollutants: a critical perspective. Environmental International, 2011; 37: 1362-1375.
- Minai-Tehrani D. and Herfatmanesh A. Biodegradation of Aliphatic and Aromatic Fractions of Heavy Crude Oil-Contaminated Soil: A Pilot Study. Journal of Biodegradation and Bioremediation, 2007; 11(2): 71-76.
- Mishra S., Jyot J., Kuhad R.C. and Lal B. Evaluation of inoculum addition to stimulate in situ bioremediation of oily sludge contaminated soil. Applied Environmental Microbiology, 2001; 67: 1675-1781.
- Nojiri H., Nam J.W., Kosaka M., Mori K.I., Takamura T., Furihata K.,Yamane H. and Omori T. Diverse oxygenations catalyzed by carbazole 1,9a-dioxygenase from Pseudomonas sp. strain CA10. Journal of Bacteriology, 1999; 181: 3105–3113.
- Obayori O.S., Ilori, M.O., Adebusoye S.A., Oyetibo G.O. and Amund O.O. Pyrene-degradation potentials of Pseudomonas species isolated from polluted tropical soils. World Journal of Microbiology and Biotechnology, 2008; 24: 2639–2646.
- Okerentugba P.O. and Ezeronye O.U. Petroleum degrading potentials of single and mixed microbial cultures isolated from rivers and refinery effluent in Nigeria. African Journal of Biotechnology, 2003; 2: 288–292.
- Okoh A.I. Biodegradation alternative in the cleanup of petroleum hydrocarbon pollutants, Biotechnology and Molecular Biology Review, 2006; 1: 38-50.
- Olajire A.A., Olujobade M. and Olabemiwo O. N-Alkane distribution in soil and water samples collected near Agbabu bitumen field of South Western Nigeria. International Journal of Environmental Studies, 2008; 65(6): 769-779.
- Olanipekun O. O., Ogunbayo A.O. and Layokun S.K. A study of the abilities of Pseudomonas aeruginosa and Pseudomonas fluorescens to degrade diesel oil. Journal of Emerging Trends in Applied Science, 2012; 3(3): 429-434.
- Sarma P., Bhattacharya D.S., Krishnan and Banwari L. Assessment of intra-species diversity among strains of Acinetobacter baumannii isolated from sites contaminated with petroleum hydrocarbons, Canadian Journal of Microbiology, 2004; 50: 405-414.
- Tao K., Liu X., Chen X., Hu X., Cao L., and Yuan X. Biodegradation of crude oil by a defined co-culture of indigenous bacterial consortium and exogenous Bacillus subtilis. Bioresour. Technol. 2017; 224, 327–332. doi: 10.1016/j. biortech. 2016.10.073
- Tri P.T. Oily wastewater treatment by hydrocarbon degradation during full scale membrane bioreactor process coupled with bioremediation. Biological activated carbon process. M.Sc. Thesis, University of Technology, Ho Chi Minh City, China. 2002.
- Vankateswaran K., Hoaki T., Kato M. and Murayama T. Microbial degradation of resins fractionated for Arabian light crude oil. Canadian Journal of Microbiology, 1995; 41: 418-424.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.