ISSN: 0973-7510
E-ISSN: 2581-690X
Silver nanoparticles (AgNPs), have been widely used as antibacterial therapy for any microorganisms that are multidrug resistance to antibiotics. Pseudomonas aeruginosa is the most common respiratory pathogen in patients with cystic fibrosis (CF), was collected from Al-Muthanna hospitals. These isolates were drug-resistant against tetracycline, chloramphenicol, erythromycin, streptomycin, azithromycin and trimethoprim, while they were sensitive to imipenem. The conventional PCR was used to screen for many different virulent genes and eventually, the only algD and plcH were detected among 11 Pseudomonas aeruginosa strains over of twenty isolates of P. aeruginosa isolated from patients with cystic fibrosis (CF) disease. The Ag/F Tio2 NPs was used as antibacterial to test the AgNPs activity against the expression of algD and plcH genes that are screened as a common complication of gene virulence in the cystic fibrosis (CF) disease. The results showed a significant effect on algD and plcH genes expression of P. aeruginosa.
Pseudomonas aeruginosa, Ag/F Tio2, Cystic Fibrosis, Nanoparticles
Silver nanoparticles are considered antimicrobial,potency agents to treat and inhibit the bacterial infection. It has been utilizedas antibacterial against multidrug resistant bacteria such as P. aeruginosa1. Silver nanoparticles have shown a strong effect on microorganisms such as bacteria, viruses and fungi compared to other minerals2. Silver nanoparticles have been used in the field of health products such as medical devices antimicrobials and applications3. Cystic fibrosis (CF) is the most common respiratory disease that is caused by a primary pathogen Pseudomonas aeruginosa in addition to Staphylococcus aureus4,5,7.
The infection with Cystic fibrosis (CF) is caused by many microorganisms such as pathogenic bacteria and fungal bacteria or fungi causes the inflammation leading to destruction of the lungs and ultimately death. Patients infected with Cystic fibrosis are more susceptible to failure in the respiratory system which is eventually cause the death6. Moreover, the mutation into conductance gene (CFTR), encoding for a membrane protein, involved in electrolytes transport. This type of mutation is leading to the destruction of epithelial cell function as the production of a result of sticky and bronchial secretions6,22,23. Pseudomonas aeruginosa is a major cause of chronic bronchial infection in cystic fibrosis8. In the CF lung, the P. aeruginosa is genetically adapt to cause infection in the lung. This modification is caused that P. aeruginosa is loss of quorum sensing (QS), which is the mechanism for productive the bacterial infection9. The P. aeruginosa is often the nosocomial bacteria as well as is multidrug resistance bacteria against many antibiotics such as fluoroquinolones, aminoglycosides and cephalosporin due to acquisition of different mechanisms for antibiotics resistance10,11.
Pseudomonas aeruginosa colonizes the lung, causing cystic fibrosis and patients infected with this disease have mucosal lesions due to the production of polysaccharide genes18. This also helps bacteria confirm bacterial biofilms that are important in bacterial infection of the lung26. Biofilm is required for both regulators which are AlgB and AlgR controlled by algD, coding for the hydrogenase GDP, which has a major role in the biosynthesis of genes18,26.
The P. aeruginosa causes many diseases upon on the virulence genes factors leading to cause the infection such as cystic fibrosis (CF). The plcH gene activity is phosphatidylcholine (PC) and sphingomyelin. This lipid makes the cellular membrane which is important in bacterial biofilm27. Many studies have been reported that the plcH gene (phospholipase C/sphingomyelinase) becomes expressed during the lung infection of P. aeruginosa28.
The aim of this study was to detect most virulence genes in P. aeruginosa isolated from patients with Cystic fibrosis (CF) and test the antibacterial activity of Ag/F Tio2 nanoparticles against these genes expressions in Pseudomonas aeruginosa.
Samples collection
In this study, 100 clinical samples of sputum were collected from patients with cystic fibrosis infection from AL-Hussein Teaching Hospital in Al Muthanna city between the period from September-2018 to April-2019.
Isolation and identification
The isolation and differentiation of the Pseudomonas aeruginosa strains were confirmed by growing the samples onto CHROMagar™ media and then were confirmed using VITEK- 2 Compact system (Biomerieux/ France)14.
DNA extraction
Bacterial genomic DNAs were extracted using the Favor Prep Tissue Genomic DNA Extraction Mini Kit (Favorgen, Taiwan) following the manufacturer’s protocol.
PCR method
The multiplex PCR was performed, after gDNA extraction from the P. aeruginosa isolates in a total volume of 25 µl using Maxime™ PCR PreMix (i-Taq) (iNtRON Biotechnology), including primers see oligo Table (1), following the manufacturer’s protocol. PCR programme for Tag was normally consisted as in Table (2).
Table (1):
Primers used in this paper.
| Oligo | Sequence | Use | PCR product (size) (bp) |
|---|---|---|---|
| YA1 | CGTCTGCCGCGAGATCGGCT | Forward to screen for algD gene | 313 |
| YA1 | GACCTCGACGGTCTTGCGGA | Reverse to screen for algD gene | |
| YA4 | GCACGTGGTCATCCTGATGC | Forward to screen for plcH gene | 608
|
| YA4 | TCCGTAGGCGTCGACGTAC | Reverse to screen for plcH gene |
Table (2):
PCR programme used for genes screen.
| Steps | Temperature | Time | Number Cycles |
|---|---|---|---|
| Initial denaturation | 94°C | 3 min | 1 |
| Denaturation | 94°C | 30 sec |
32 |
| Annealing | 55°C | 30 sec | |
| Extension | 72°C | 1 min per kb | |
| Final extension | 72°C | 10 min | 1 |
| Hold | 12°C | ∞ |
Once the PCR reactions were completed, separated on a 0.8% agarose gel at 100V for 25 min and then the DNA samples were visualised on a UV transilluminator.
Nanoparticles preparation
The Ag/F Tio2 nanoparticles were collected from College of Science for Women-University of Babylon at final concentration of 10 mg/ml and we diluted to 1 mg/ml as final concentration with bacterial broth.
Antibacterial activity of Ag/F Tio2 nanoparticles
The antimicrobial activity of Ag/F Tio2 was evaluated by growing the suspension of culturing bacteria to a final concentration of 1 mg/ml of Ag/F Tio2. Subsequently, the gDNA was extraction from the samples before and after nanoparticles treatment then the PCR was performed to test the effect of the nanoparticles of both algD and plcH genes expression.
Antibiotic Susceptibility Test
The antibiotic susceptibility test for P. aeruginosa was done using disc diffusion on Mueller-Hinton agar (CM0337-OXOID), following15. Briefly, the standard of P. aeruginosa in oculum of 1.5*108 cell was cultured on Mueller Hintonagar. Subsequently, the antibiotics discs were placed on contaminated plates. Finally,the plates were incubated at 37°C for 24-48 h, then the diameter of the inhibition zone was measured by mm15.
Statistical Methods
The antibiotic resistance patterns were tested by the measured using ANOVA, GraphPad Prism software.
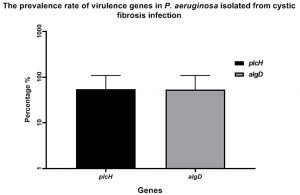
In the study, clinical and resistant isolates of P. aeruginosa were isolated from cystic fibrosis (CF)and screened for different virulence genes that are important in causing P. aeruginosa pathogenesis. One hundred clinical sputum were collected, followed by culturing onto CHROMagar™ media to isolated P. aeruginosa strains. Eventually, twenty strains were isolated and confirmed by using VITEK- 2 Compact6system. Interestingly, the PCR screen was shown the detection of both algD and plcH genes at prevalence rate 55% of P. aeruginosa strains isolated from cystic fibrosis (CF) Fig. (1).
Fig. 1. The prevalence rate of virulence gene of P. aeruginosa isolates from cystic fibrosis (CF). The gene algD and plcH were detected in P. aeruginosa strains at a rate of 55%.
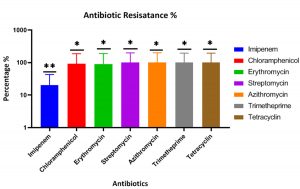
The antibiotics sensitivity for the P. aeruginosa isolates against different antibiotics was shown the highest resistance percentage for tetracycline, chloramphenicol, erythromycin, streptomycin, azithromycin, trimethoprim, while the lowest resistance against the imipenem at 20% at the p-value of 0.0212 (Fig. 2).
Fig. 2. The percentage of antibiotic resistance of P. aeruginosa strains isolated from cystic fibrosis (CF) in al Muthanna hospitals shows the highest percentage of the resistance for all antibiotics were tested, while the strains were shown the lowest resistance level against imipenem. The p-value of antibiotics resistance is 0.0212 that measured using ANOVA, GraphPad Prism software. **: the population means are significantly different at the p-value= 0.05 level.
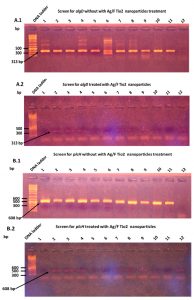
The results of PCR amplification to specific algD and plcH genes was detected in all P. aeruginosa isolates from cystic fibrosis (CF) infections. The PCR was gave the expected product of 313 bp for algD gene, Fig. (3-A1), and the PCR amplification showed the expected product of 608 bp (Fig. 3-B1).
Fig. 3. PCR amplification of both algD and plcH genes P. aeruginosa isolated from patients with Cystic Fibrosis. before and after Ag/F Tio2 treatment. A.1: PCR amplification of algD, showing amplified the predicted product of 313 bp. A.2: The algD gene amplification after Ag/F Tio2 treatment showing the Ag/F Tio2 nanoparticles effect on algD gene amplification. B.1: The plcH gene amplification showing amplified the predicted product of 608 bp. B.2 The plcH gene amplification after Ag/F Tio2 treatment showing the Ag/F Tio2 nanoparticles effect on plcH gene amplification. DNA ladder: molecular weight: 100–10000 bp.
The P. aeruginosa strains that were given positive results for algD and plcH genes detection were then treated with Ag/F Tio2 nanoparticles. The P. aeruginosa strains were grown up with Ag/F Tio2 nanoparticles using brain heart infusion broth and incubated at 37ᵒC for 24 h. The gDNA was extracted from treated samples. The PCR amplification showed that disappearance of gene expression for both algD and plcH genes (Figs. 3-A2 & 3-B2). This result suggested that the Ag/F Tio2 nanoparticles have an effect on down-regulation of both algD and plcH genes in the P. aeruginosa.
Pseudomonas aeruginosa is a common complication for many infections such as cystic fibrosis (CF) and wound, associated with resistance for multi-drug antibiotics resistance, leading to the increase of the rate of both morbidity and mortality16,19.
Pseudomonas aeruginosa is Gram-negative bacteria have been recently treated with AgNPs showing a strong effect on gene expression by up-regulated 27 genes and 32 down-regulated genes17. The actual mechanism of AgNPs is the interference with the function of the bacterial cell membrane leading to generate reactive oxygen species (ROS). Moreover, the AgNPs have a similar role to silver ions in the regulation of the membrane transporter proteins. These mechanisms lead to an effect on bacterial growth and inhibition of multi drugs resistance bacteria17. Likewise, the silver nanoparticles have been tested against different bacteria such as E. coli, S. aureus methicillin-resistant, K. pneumonia and P. aeruginosa showing a promising activity but the activity against P. aeruginosa was the highest at a rate of 90% compared with others bacteria21.
The PCR screen results showed the detection for both algD and plcH genes and this is agreed with others studied for Pseudomonas aeruginosa strains isolated from the lungs with cystic fibrosis disease24. The transcription of algD gene, which encodes to GDP-mannose dehydrogenase, causing for production of exopolysaccharide alginate that is important in the infection of Pseudomonas aeruginosa in the lung18. However, the plcH gene expression is responsible for biofilm formation due to making lipid in the bacterial cell membrane and the mutant of this gene have been shown a defective in colonizing in the lung of a model animal with no biofilm formation20.
In conclusion, this study has demonstrated that the Ag/F Tio2 nanoparticles can affect the expression of the most important virulence genes (algD and plcH) in P. aeruginosa during the lung infection. Our results suggest that the silver nanoparticles can be used as alternative antibacterial therapy against cystic fibrosis disease.
ACKNOWLEDGMENTS
We would like to express our heartfelt thanks to Professor Ayad Alkaim for providing nanoparticles particles to study the effect on gene expression.
CONFLICT OF INTEREST
The authors declare that there is no conflicts of interest.
AUTHORS’ CONTRIBUTION
All authors designed and performed the experiments. YA analyzed the data, YA and MQ wrote the manuscript. All authors read and approved the manuscript.
FUNDING
None.
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript and/or the Supplementary Files.
- Salomoni R, Leo P, Montemor A, Rinaldi B, Rodrigues M. Antibacterial effect of silver nanoparticles in Pseudomonas aeruginosa. Nanotechnol Sci Appl. 2017;10:115.
Crossref - Sahayaraj K, Rajesh S. Bionanoparticles: synthesis and antimicrobial applications. Science against Microbial Pathogens: Communicating Current Research and Technological Advances. 2011;23:228-244.
- Hossain A, Hong X, Ibrahim E, et al. Green Synthesis of Silver Nanoparticles with Culture Supernatant of a Bacterium Pseudomonas rhodesiae and their antibacterial activity against soft rot pathogen Dickeya dadantii. Molecules. 2019;24:2303.
Crossref - Speert DP, Campbell ME, Henry DA, et al. Epidemiology of Pseudomonas aeruginosa in cystic fibrosis in British Columbia, Canada. Am J Respir Crit Care Med. 2002;166:988-993.
Crossref - de Dios Caballero J, del Campo R, Tato M, et al. Microbiological diagnostic procedures for respiratory cystic fibrosis samples in Spain: towards standard of care practices. BMC Microbiology. 2014;14:335.
Crossref - Gilligan PH. Infections in patients with cystic fibrosis: diagnostic microbiology update. Clin Lab Med. 2014;34:197-217.
Crossref - Vongthilath R, Thiriez BR, Dehillotte C, et al. Clinical and microbiological characteristics of cystic fibrosis adults never colonized by Pseudomonas aeruginosa: Analysis of the French CF registry. PloS One. 2019;14:e0210201.
Crossref - Strateva T, Yordanov D. Pseudomonas aeruginosa–a phenomenon of bacterial resistance. Indian J Med Microbiol. 2009;58:1133-1148.
Crossref - Waters CM, Goldberg JB. Pseudomonas aeruginosa in cystic fibrosis: A chronic cheater. Proceedings of the National Academy of Sciences. 2019;116:6525-6527.
Crossref - Sonbol FI, Khalil MAEF, Mohamed AB, Ali SS. Correlation between antibiotic resistance and virulence of Pseudomonas aeruginosa clinical isolates. Turk J Med Sci. 2015;45:568-577.
Crossref - Rocha AJ, Barsottini MRdO, Rocha RR, et al. Pseudomonas aeruginosa: Virulence Factors and Antibiotic Resistance Genes. Braz Arch Biol Technol. 2019;62.
Crossref - Jalali S, Allafchian A, Banifatemi S, Tamai IA. The antibacterial properties of Ag/TiO2 nanoparticles embedded in silane sol–gel matrix. J Taiwan Inst Chem Eng. 2016;66:357-362.
Crossref - Stehling EG, Leite DS, Silveira WD. Molecular typing and biological characteristics of Pseudomonas aeruginosa isolated from cystic fibrosis patients in Brazil. Braz J Infect Dis. 2010;14:462-467.
Crossref - MacFaddin JF. Biochemical tests for identification of medical bacteria (3rded.), Lippincott Williams and Wilkins, USA, 2000.
- Benson HJ. Antimicrobial sensitivity testing: The Kirby-Bauer method. In: Benson HJ, editor. Microbiological Application: Laboratory Manual in General Microbiology. 7th ed. Boston: McGraw Hill, 1998;139-41.
- Ijaz M, Siddique AB, Rasool MH, Shafique M. Frequency of multi drug resistant Pseudomonas aeruginosa in different wound types of hospitalized patients. Pak J Pharm Sci. 2019;32.
- Yan X, He B, Liu L, et al. Antibacterial mechanism of silver nanoparticles in Pseudomonas aeruginosa: proteomics approach. Metallomics. 2018;10:557-564.
Crossref - Wozniak DJ, Ohman D. Transcriptional analysis of the Pseudomonas aeruginosa genes algR, algB, and algD reveals a hierarchy of alginate gene expression which is modulated by algT. Journal of Bacteriology, 1994;176:6007-6014.
Crossref - Faraji F, Mahzounieh M, Ebrahimi A, Fallah F, Teymournejad O, Lajevardi B. Molecular detection of virulence genes in Pseudomonas aeruginosa isolated from children with Cystic Fibrosis and burn wounds in Iran. Microb Pathog. 2016;99:1-4.
Crossref - Jackson AA, Gross MJ, Daniels EF, et al. Anr and its activation by PlcH activity in Pseudomonas aeruginosa host colonization and virulence. J Bacteriol. 2013;195:3093-3104.
Crossref - Eid M, Araby E. Bactericidal effect of poly (acrylamide/itaconic acid)–silver nanoparticles synthesized by gamma irradiation against Pseudomonas aeruginosa. Appl Biochem Biotechnol. 2013;171:469-487.
Crossref - Goss CH, Ratjen F. Update in cystic fibrosis 2012. Am J Respir Crit Care Med. 2013;187:915-919.
Crossref - Knapp EA, Fink AK, Goss CH, et al. The Cystic Fibrosis Foundation Patient Registry. Design and methods of a national observational disease registry. Ann Am Thorac Soc. 2016;13:1173-1179.
Crossref - Mitov I, Strateva T, Markova B. Prevalence of virulence genes among bulgarian nosocomial and cystic fibrosis isolates of Pseudomonas aeruginosa. Braz J Microbiol. 2010;41:588-595.
Crossref - Snyder A, Vasil AI, Zajdowicz SL, Wilson ZR, Vasil ML. Role of the Pseudomonas aeruginosa PlcH Tat signal peptide in protein secretion, transcription, and cross-species Tat secretion system compatibility. Journal of Bacteriology. 2006;188:1762-1774.
Crossref - Hoffmann N, Rasmussen TB, Jensen P, et al. Novel mouse model of chronic Pseudomonas aeruginosa lung infection mimicking cystic fibrosis. Infect Immun. 2005;73:2504-2514.
Crossref - Lopez DJ, Collado MI, Ibarguren M, et al. Multiple phospholipid substrates of phospholipase C/sphingomyelinase HR2 from Pseudomonas aeruginosa. Chem Phys Lipids. 2011;164:78-82.
Crossref - Wiener-Kronish JP, Sakuma T, Kudoh I, et al. Alveolar epithelial injury and pleural empyema in acute P. aeruginosa pneumonia in anesthetized rabbits. J Appl Physiol. 1993;75:1661-1669.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.