ISSN: 0973-7510
E-ISSN: 2581-690X
Urinary tract infections (UTIs) remain one of the most common bacterial infections worldwide, where timely identification of causative pathogens and their antibiotic susceptibility is critical for guiding therapy. Conventional culture-based methods are reliable but slow, creating an urgent need for rapid, point-of-care (POC) diagnostic solutions. This study aimed to develop and validate a novel Alamar Blue-based system capable of simultaneous Gram classification and antibiotic susceptibility testing (AST) directly from urine samples. The system utilizes Alamar Blue (resazurin), a redox indicator that shifts from violet (oxidized) to pink (reduced) in response to bacterial metabolic activity. Gram differentiation was achieved using two selective antibiotic marker vials: vancomycin/linezolid for Gram-positive and aztreonam/polymyxin B for Gram-negative bacteria. New Gram differentiation standardization (GDS) was performed using the CLSI MIC breakpoint method, ensuring clinical relevance. To improve accuracy, urine artifacts were removed through a filtration approach prior to analysis. Validation on ATCC reference strains demonstrated 100% agreement in Gram classification and 97% similarity with the conventional Kirby–Bauer disc diffusion method. On real urine samples (n = 25), the newly developed GDS predicted 24% of samples as negative, showing 100% concordance with culture-based findings. Furthermore, 16% of samples were identified as true positives, also showing 100% agreement with conventional laboratory reports. AST outcomes produced clear colorimetric readouts within five hours without requiring complex infrastructure. This Alamar Blue-based POC platform provides a rapid, user-friendly, and cost-effective alternative to conventional diagnostics, enabling both Gram differentiation and AST from urine specimens in resource-limited healthcare settings.
Resazurin Reagent, Gram Classification, Point-of-Care Test, Rapid Urinary Tract Diagnosis, Antibiotic Sensitivity Test
Urinary tract infections (UTIs) are among the most prevalent infectious diseases, estimated 150 million global cases annually. UTIs constitute a significant burden on public health systems, particularly in developing countries, due to their high recurrence rate and potential for serious complications.1,2 Anatomical and physiological differences render women substantially more susceptible to UTIs than males, further amplifying the clinical and epidemiological impact of these infections.3,4 Clinically, UTIs may present with a spectrum of symptoms ranging from cystitis and bacteriuria to more severe manifestations such as pyelonephritis.5 In the absence of timely and appropriate intervention, the infection can ascend from the lower to the upper urinary tract, potentially leading to renal damage and systemic complications.6 The microbial etiology of UTIs is well established. A multicentric surveillance study analysing 2,459 urine specimens revealed 250 culture-positive cases, of which 94% were caused by Gram-negative bacilli and only 6% by Gram-positive cocci.7 The distribution of pathogens included E. coli (68.3%), Klebsiella spp. (17.6%), Proteus spp. (3.2%), Acinetobacter spp. (1.2%), and Enterococcus spp. (5.6%), reflecting the dominance of Gram-negative organisms in UTI pathogenesis.8 These findings emphasize the need for rapid, accurate diagnostic systems for early detection and appropriate antimicrobial therapy, especially in the face of rising antimicrobial resistance (AMR) and the clinical complexity associated with delayed treatment.9
Despite the well established burden of urinary tract infections (UTIs), traditional diagnostic workflows for pathogen identification and antibiotic susceptibility testing (AST) remain heavily reliant on centralized laboratory infrastructure, particularly in tertiary care settings. The conventional diagnostic process typically involves a sequence of steps, including urine centrifugation, wet mount preparation, Gram staining, semi-quantitative culture-based colony counting, biochemical characterization, and AST via the Kirby-Bauer disc diffusion method.10 This multi-step protocol often spans up to 72 hours, emphasizing comprehensive diagnosis but frequently delaying timely therapeutic intervention. Public Health Centres often lack essential infrastructure, including microbiological laboratories, experienced technicians, basic diagnostic reagents, and even functional microscopy units. These limitations severely hinder the implementation of culture-based diagnostics and AST in primary health care settings.11 These challenges underscore the urgent need for accessible, rapid, and reliable point-of-care (POC) diagnostic systems specifically tailored to the constraints of primary healthcare settings in UTI diagnosis and drug therapy.
To overcome diagnostic delays and infrastructural constraints of conventional urine culture and AST, point-of-care (POC) technologies have been developed to provide rapid, user-friendly alternatives for UTI diagnosis.12,13 Dipstick-based assays are widely used due to their affordability, but their reliance on surrogate markers like leukocyte esterase and nitrites results in suboptimal sensitivity and specificity, with false positives in non-infectious conditions and false negatives for pathogens such as Pseudomonas aeruginosa, which cannot reduce nitrate to nitrite.14 While tertiary hospitals can compensate through extended cultures, primary healthcare centres (PHCs) lack such capacity, leading to underdiagnosis or empirical treatments. Recent innovations include paper-based analytical devices detecting E. coli in six hours from ~20 µL of urine (104 CFU/mL), fluorescent immunoassays with higher sensitivity (240 CFU/mL in 30 minutes), and digital dipsticks reporting within eight hours at 253 CFU/mL.15 Fluorescent sensors for detecting Gram-negative bacteria also exist, but require large sample volumes and high bacterial loads.16 Despite rapid results, all are pathogen-specific, impractical for diverse UTI etiologies, and lack integrated AST. A UTI diagnostic kit includes combined pathogen detection with AST but excludes Gram-positives, limiting PHC use. Future POC platforms must ensure broad pathogen coverage, Gram differentiation, simple colorimetric readouts, and integrated AST for effective UTI management in PHC.
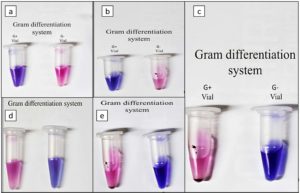
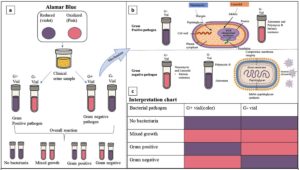
This research article marks a significant step toward developing a point-of-care diagnostic system for urinary tract infections (UTIs), particularly suited for resource-limited primary healthcare settings. A novel Alamar Blue-based colorimetric assay was developed and validated for gram differentiation and AST detection. This integrated approach allows for rapid Gram classification and antibiotic susceptibility testing (AST) within just 5 hours using urine samples. The newly developed Alamar Blue-based diagnostic system enables simultaneous Gram classification and antibiotic susceptibility testing (AST) from a single urine sample. Alamar Blue (resazurin), a redox-sensitive dye, is violet in its oxidized state but turns pink upon reduction by metabolically active bacteria, thus indicating viability (Figure 1a). The Gram differentiation system (GDS) employs two vials containing selective antibiotic markers. The Gram-positive vial includes vancomycin and linezolid, which inhibit Gram-positive bacteria by disrupting cell wall synthesis and protein translation, respectively. Conversely, the Gram-negative vial contains aztreonam and polymyxin B, which target the outer membrane and peptidoglycan synthesis of Gram-negative organisms (Figure 1b). When a urine sample contains Gram-positive bacteria, the Gram-positive vial remains violet due to inhibition, while the Gram-negative vial turns pink from metabolic activity. In Gram-negative infections, the opposite occurs. Absence of bacteriuria yields no color change in either vial, whereas polymicrobial samples produce pink coloration in both, suggesting mixed growth and requiring confirmatory laboratory evaluation (Figure 1c).
Figure 1. a. Alamar blue-based gram differentiation assay, b. Mechanism basis of gram categorization using antibiotics, c. Interpretation chart for gram differentiation results
Following Gram differentiation, AST is performed using individual antibiotic vials, each prepared at CLSI breakpoint concentrations. The system covers six antibiotics for Gram-positive and eight for Gram-negative pathogens. In each vial, a violet color indicates susceptibility, while a pink shift denotes resistance. This rapid, visual readout bypasses the need for sophisticated equipment and aligns with established Alamar Blue antimicrobial assays. By integrating Gram classification with AST, this point-of-care platform provides an accurate, user-friendly, and resource-efficient approach for guiding targeted therapy, particularly in decentralized or resource-limited clinical settings.
All American Type Culture Collection (ATCC) bacterial strains used in this study were collected from the Department of Microbiology, Aarupadai Veedu Medical College and Hospital, Puducherry, maintained for antibiotic sensitivity testing. The ATCC strains used include Escherichia coli (ATCC 25922), Staphylococcus aureus (ATCC 25923), Pseudomonas aeruginosa (ATCC 27853), Klebsiella pneumoniae (ATCC 13883), and Enterococcus faecalis (ATCC 29212). The Resazurin reagent was purchased from HiMedia. Whatman number 2, diameter 90 mm, was purchased from Cytiva.
New GDS development and standardization
investigations with vancomycin and aztreonam
ATCC strains of Staphylococcus aureus (ATCC 25923), Escherichia coli (ATCC 25922), Klebsiella pneumoniae (ATCC 13883), Pseudomonas aeruginosa (ATCC 27853), and Enterococcus faecalis (ATCC 29212) were adjusted to a population of 106 CFU/mL. For each bacterium, two vials were prepared for the assay. The first vial was added with 250 µL of vancomycin solution (30 µg/mL), and the second vial was added with 250 µL of aztreonam solution (30 µg/mL). Each vial was subsequently inoculated with 250 µL of the population-adjusted bacterial suspension prepared in Brain Heart Infusion (BHI) broth. The vials were then incubated at 37 °C for 5 hours. Following incubation, 75 µL of resazurin reagent was added to each vial, and color changes were monitored.
Gram’s categorization using drug combinations
ATCC strains of Staphylococcus aureus (ATCC 25923), Escherichia coli (ATCC 25922), Klebsiella pneumoniae (ATCC 13883), Pseudomonas aeruginosa (ATCC 27853), and Enterococcus faecalis (ATCC 29212) were adjusted to a population of 106 CFU/mL. For each bacterium, two vials were prepared for the assay. The first vial was added with 250 µL of vancomycin solution (30 µg/mL) and linezolid, and the second vial was added with 250 µL of aztreonam solution (30 µg/mL) and polymyxin B. Each vial was subsequently inoculated with 250 µL of the population-adjusted bacterial suspension prepared in Brain Heart Infusion (BHI) broth. The vials were then incubated at 37 °C for 5 hours. Following incubation, 75 µL of resazurin reagent was added to each vial, and color changes were monitored.
Drug combination concentration optimization for E. coli
The ATCC strains of E. coli were adjusted to a population of 106 CFU/mL. A series of vials prepared with different Vancomycin concentrations, like 4 µg/mL + Linezolid, 8 µg/mL + Linezolid, 12 µg/mL + Linezolid, 16 µg/mL + Linezolid, 20 µg/mL + Linezolid. The tubes were inoculated with 250 µL of population-adjusted BHI broth and incubated for 5 hours. After incubation, the tubes were supplemented with 75 µL resazurin reagent. 20 minutes were kept undisturbed at room temperature, and the color changes were observed.
Gram’s categorization and AST detection
3 ml bacterial suspensions of ATCC strains – Staphylococcus aureus was taken in a tube, and the density was adjusted to 0.5 McFarland standard. Subsequently transferred to a tube containing 11 mL of BHI broth. The mixtures were thoroughly homogenized. In a 128-pocketed tray, 250 µL of incubated BHI broth was transferred into 1.5 mL vials containing 250 µL of various antibiotics-saline mixture (AST), along with the separate vials designated for Gram’s categorization (GDS). The vials were incubated at 37 °C for 5 hours. After incubation, 75 µL of resazurin solution was added to each vial. The vials were incubated for an additional 15 minutes, after which the color change was observed both in the Gram’s categorization vials and the antibiotic vials. The same ATCC strains were tested using the Kirby-Bauer disc diffusion method, and the results were compared. This procedure was similarly performed with other bacterial strains, including Escherichia coli (ATCC 25922), Klebsiella pneumoniae (ATCC 13883), Pseudomonas aeruginosa (ATCC 27853), and Enterococcus faecalis (ATCC 29212).
Filtration-based approaches to remove pus cells and other artifacts
Urine samples with significant pyuria were collected. A 3 ml urine sample was taken in a test tube and centrifuged at 1500 rpm for 5 minutes. The wet mount of samples prepared from the sediment. Pus cells and other artifact distributions were noted. The same samples 3ml filtered through the Whatman no 2 filter paper, and the filtrate was taken out and centrifuged at 1500 rpm for 5 minutes, and the pus cells and other artifacts were noted.
Gram’s identification system screening and AST detection in urine
A filtered 3 mL urine sample was transferred into a test tube, adding 13 mL of BHI, and the tube was left undisturbed for 15 minutes to promote bacterial enrichment. After the enrichment period, the solution was vortexed briefly to ensure uniform mixing, and 250 µL of the enriched sample was transferred to pre-prepared Gram’s differentiation and AST (antibiotic susceptibility testing) detection vials. All vials were then incubated for 5 hours at 37 °C. After incubation, 75 µL of resazurin indicator was added to each vial, mixed gently, and left undisturbed for 15 minutes to allow color development. The results were recorded after 15 minutes.
New GDS development and standardization
investigations with vancomycin and aztreonam
The initial studies utilizing vancomycin (30 µg/mL) and aztreonam (30 µg/mL) demonstrated effective categorization of Gram-positive and Gram-negative bacteria in four out of the five selected ATCC strains. However, Escherichia coli exhibited incorrect color reactions, indicating potential variability in its response. In all other selected ATCC strains, intrinsic resistance mechanisms were clearly observed. Staphylococcus aureus and Enterococcus faecalis, both Gram-positive organisms, were found sensitive to vancomycin, resulting in a violet color in the vancomycin-containing vials after 5 hours of incubation. Conversely, vials containing aztreonam for these strains changed to pink, indicating intrinsic resistance. Similarly, all Gram-negative pathogens except E. coli such as Pseudomonas aeruginosa and Klebsiella pneumoniae were sensitive to aztreonam, maintaining a violet color in Gram-negative vials. These Gram-negative bacteria showed intrinsic resistance to vancomycin, as evidenced by the color change in the vancomycin-containing vials to pink. The variations in the color appearance for E. coli failed the system, leading to the decision to standardize it with a combination of drugs (Table 1).
Table (1):
New GDS development and standardization
| Attempts with different antibiotic concentrations | Antibiotics and concen. | Organisms | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Staphylococcus aureus | Enterococcus faecalis |
Escherichia coli |
Pseudomonas aeruginosa |
Klebsiella pneumoniae | |||||||
| Color | Interpretation | Color | Interpretation | Color | Interpretation | Color | Interpretation | Color | Interpretation | ||
| Vancomycin and Aztreonam | Gram-positive vial (Vancomycin 30 µg/ml) | Gram-positive pathogen | Gram-positive pathogen | Gram-positive pathogen (wrong interpretation) | Gram-negative pathogen | Gram-negative pathogen | |||||
| Gram-negative vial (Aztreonam 30 µg/ml) | |||||||||||
| Drug combinations | Gram-positive vial Vancomycin 30 µg/ml + Linezolid | Gram-positive pathogen | Gram-positive pathogen | Sample without bacteria (wrong interpretation) | Gram-negative pathogen | Gram-negative pathogen | |||||
| Gram-negative vial Aztreonam 30 µg/ml + Polymyxin B | |||||||||||
| After Vancomycin optimization | Gram-positive vial Vancomycin 8µg/ml + Linezolid | Gram-positive pathogen | Gram-positive pathogen | Gram-negative pathogen | Gram-negative pathogen | Gram-negative pathogen | |||||
| Gram-negative vial Aztreonam 30 µg/ml + Polymyxin B | |||||||||||
Gram’s categorization using drug combinations
The initial studies utilizing vancomycin (30 µg/mL) and aztreonam (30 µg/mL) demonstrated effective categorization of Gram-positive and Gram-negative bacteria in four out of the five selected ATCC strains. However, Escherichia coli exhibited incorrect color reactions, indicating potential variability in its response. The variations in the color appearance of E. coli resulted in the failure of the system, leading to the decision to standardize it with a combination of drugs. Following a thorough review of the literature, combinations of vancomycin with linezolid and aztreonam with polymyxin B were selected for further optimization studies. Based on these investigations, all bacterial strains except E. coli demonstrated consistent results. For E. coli, the color reactions with both drug combinations appeared as violet/violet, resolving the issue of aztreonam resistance with the addition of polymyxin B. However, E. coli remained sensitive to the vancomycin-linezolid combination. To address this challenge in subsequent studies, it was decided to optimize the concentration of vancomycin to improve the system’s accuracy for E. coli (Table 1).
Vancomycin concentration optimization for E. coli
Concentration optimization studies were conducted using a range of vancomycin concentrations (4 µg/mL to 28 µg/mL) combined with linezolid. It was observed that E. coli became sensitive to vancomycin at concentrations less than 8 µg/mL when combined with linezolid. At this optimized concentration, all selected bacterial strains demonstrated correct color reactions. Subsequent validation confirmed that the Gram differentiation system worked effectively at this concentration, providing accurate results for all tested strains (Table 1 and Figure 2).
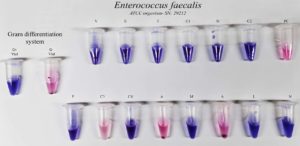
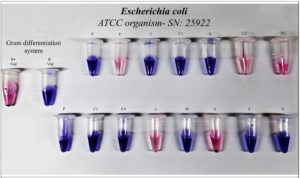
Figure 2. (a). Staphylococcus aureus (ATCC 25923), (b). Enterococcus faecalis (ATCC 25212), (c). Escherichia coli (ATCC 25922), (d). Klebsiella pneumoniae (ATCC 13883), (e). Pseudomonas aeruginosa (ATCC 27853)
Alamar blue-based GDS and AST in ATCC
The newly developed Gram differentiation system and AST detection using the Alamar Blue assay were effectively categorized the five selected ATCC strains into their appropriate Gram categories and provided AST pattern. Validation of the Gram’s identification system on ATCC strains showed 100% concordance with the Gram categories of the selected strains. Additionally, a 97% similarity was observed in AST results when compared to the disc diffusion method. For detecting AST patterns in Gram-positive bacteria, the antibiotics tested included Vancomycin (30 µg/mL) (V), Erythromycin (15 µg/mL) (E), Tetracycline (10 µg/mL) (T), Ciprofloxacin (30 µg/mL) (C1), Nitrofurantoin (100 µg/mL) (N), and Chloramphenicol (5 µg/mL) (C2). For Gram-negative bacteria, the antibiotics screened were Piperacillin/Tazobactam (100/10 µg/mL) (P), Ceftazidime (30 µg/mL) (C3), Cefepime (30 µg/mL) (C4), Aztreonam (30 µg/mL) (A), Meropenem (18 µg/mL) (M), Amikacin (30 µg/mL) (A), Levofloxacin (5 µg/mL) (L), and Nitrofurantoin (100 µg/mL) (N).
In the case of Staphylococcus aureus (ATCC 25923), the Gram differentiation assay produced a clear violet color in the Gram-positive vial and a pink color in the Gram-negative vial after 5 hours of incubation. This confirmed the organism’s Gram-positive. Based on this categorization, an appropriate antibiotic panel for Gram-positive bacteria was selected for antibiotic susceptibility testing (AST) interpretation. The strain was tested against six antibiotics: Vancomycin (30 µg/mL), Erythromycin (15 µg/mL), Tetracycline (10 µg/mL), Ciprofloxacin (30 µg/mL), Nitrofurantoin (100 µg/mL), and Chloramphenicol (5 µg/mL). S. aureus showed sensitivity to all six antibiotics tested (Figure 3).
Similarly, Enterococcus faecalis (ATCC 29212) showed a violet color in the Gram-positive vial and a pink color in the Gram-negative vial after 5 hours of incubation, confirming its Gram-positive nature. Following this categorization, the same Gram-positive antibiotic panel was used for AST. The strain demonstrated sensitivity to all six antibiotics tested: Vancomycin (30 µg/mL), Erythromycin (15 µg/mL), Tetracycline (10 µg/mL), Ciprofloxacin (30 µg/mL), Nitrofurantoin (100 µg/mL), and Chloramphenicol (5 µg/mL) (Figure 4).
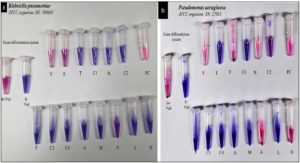
For the tested Gram-negative ATCC strains, the Gram differentiation assay resulted in a pink color in the Gram-positive vial and a violet color in the Gram-negative vial after 5 hours of incubation, confirming the Gram-negative identity of the bacteria. Upon accurate Gram classification, a relevant Gram-negative antibiotic panel was used for sensitivity profiling. Klebsiella pneumoniae (ATCC strain) demonstrated sensitivity to Piperacillin/ Tazobactam (100/10 µg/mL), Ceftazidime (30 µg/mL), Cefepime (30 µg/mL), Meropenem (18 µg/mL), Levofloxacin (5 µg/mL), Nitrofurantoin (100 µg/mL), and Aztreonam (30 µg/mL), but showed resistance to Amikacin (30 µg/mL) (Figure 5a). Pseudomonas aeruginosa (ATCC strain) demonstrated sensitivity to Piperacillin/ Tazobactam (100/10 µg/mL), Ceftazidime (30 µg/ mL), Cefepime (30 µg/mL), Meropenem (18 µg/ mL), Levofloxacin (5 µg/mL), and Aztreonam (30 µg/mL). However, the strain exhibited resistance to Amikacin (30 µg/mL) and Nitrofurantoin (100 µg/mL) (Figure 5b).In the case of Escherichia coli (ATCC-25922), the bacterium exhibited sensitivity to Piperacillin/Tazobactam (100/10 µg/mL), Ceftazidime (30 µg/mL), Cefepime (30 µg/mL), Meropenem (18 µg/mL), Levofloxacin (5 µg/mL), and Nitrofurantoin (100 µg/mL), while showing resistance to Amikacin (30 µg/mL) and Aztreonam (30 µg/mL)(Figure 6).
Figure 5. Gram’s differentiation system and AST in ATCC strains (a) Klebsiella pneumoniae and (b) Pseudomonas aeruginosa
In this method, the clinical sample is simultaneously introduced into both the Gram Differentiation System (GDS) and the Antibiotic Susceptibility Testing (AST) panels, which include antibiotics for both Gram-positive and Gram-negative organisms. Within 5 hours of incubation, distinct color changes appear in both systems. The color pattern in the GDS allows for rapid identification of the organism’s Gram nature, which in turn guides the immediate selection of the appropriate antibiotic panel for interpretation. This synchronized workflow facilitates early, targeted antibiotic decisions and sets this method apart from conventional point-of-care tests by providing integrated Gram identification and susceptibility profiling in a single step.
Filtration-based approaches to remove pus cells and other artifacts
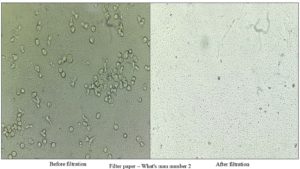
The investigations using Whatman No. 2 filter paper for urine samples demonstrated remarkable efficacy in removing pus cells and other artifacts from the samples. Among the samples subjected to filtration, all were found to have complete removal of pus cells upon further analysis. Importantly, the filtration process selectively removed pus cells and artifacts, while retaining bacteria in the filtrate, facilitating subsequent studies (Figure 7). The performance of the Whatman No. 2 filter paper highlights its potential in the differential detection of pyuria (presence of pus cells) and bacteriuria (presence of bacteria) in urine samples. Conjugating the filter paper with diazonium salts and introducing resazurin reagent to the filtrate enabled the simultaneous detection of pyuria and bacteriuria in the same sample.
Figure 7. Changes observed in a urine sample with significant pyuria (50 pus cells/high-power field) after filtration
Gram’s categorization and AST detection in Urine for POC
Out of the 25 clinical urine samples tested, both the conventional laboratory method and the newly developed Gram Differentiation System (GDS) confirmed the absence of bacteria in 16 samples. The GDS reported a Violet/Violet color pattern indicating no bacterial presence for samples U3245, U3252, U3254, U3259, U3246, U3248, U3251, U3247, U3278, U3281, U2857, U2862, U2867, U2874, U2906, and U2934 (Table 2).
Table (2):
Gram’s categorization and AST detection in Urine for POC Real urine sample studies
| Gram’s categorization system | AVMC lab findings | |||||
|---|---|---|---|---|---|---|
| No. | Sample no | Time | Gram-positive vial (V-violet) (P-Pink) | Gram-negative vial (V-violet) (P-Pink) | interference | |
| 1 | U3245 | 5-7 | V | V | No growth | No growth |
| 2 | U3252 | 5-7 | V | V | No growth | No growth |
| 3 | U3254 | 5-7 | V | V | No growth | No growth |
| 4 | U3249 | 5-7 | V | V | No growth | No growth |
| 5 | U3246 | 5-7 | V | V | No growth | No growth |
| 6 | U3248 | 5-7 | V | V | No growth | No growth |
| 7 | U3250 | 5-7 | P | P | Mixed growth | Mixed growth |
| 8 | U3255 | 08-10 | P | P | Mixed growth | Mixed gram-positive and gram-negative growth |
| 9 | U3251 | 5-7 | V | V | No growth | No growth |
| 10 | U3253 | 5-7 | P | V | Gram-negative bacteria | Gram-positive and gram-negative bacteria |
| 11 | U3247 | 5-7 | V | V | No growth | No growth |
| 12 | U3264 | 5-7 | V | P | Gram-positive bacteria | Gram-positive bacteria (contaminant) |
| 13 | U3266 | 5-7 | P | P | Mixed growth | Gram-negative bacilli with yeast |
| 14 | U3274 | 5-7 | P | V | Gram-negative bacteria | E. coli |
| 15 | U3284 | 5-7 | P | V | Gram-negative bacteria | E. coli |
| 16 | U3278 | 08-10 | V | V | No growth | No growth |
| 17 | U3281 | 5-7 | V | V | No growth | No growth |
| 18 | U3266R | 5-7 | P | V | Gram-negative bacteria | E. coli |
| 19 | U3298 | 5-7 | P | P | Mixed growth | Gram-negative bacilli with Acinetobacter |
| 20 | U2857 | 5-7 | V | V | No growth | No growth |
| 21 | U2862 | 5-7 | V | V | No growth | No growth |
| 22 | U2867 | 5-7 | V | V | No growth | No growth |
| 23 | U2874 | 5-7 | V | V | No growth | No growth |
| 24 | U2906 | 5-7 | V | V | No growth | No growth |
| 25 | U2934 | 5-7 | V | V | No growth | No growth |
Among the remaining nine samples, conventional laboratory investigations identified four as true positives with significant bacteriuria, while the other five were considered contaminated, likely due to improper sample collection. Specifically, samples U3274, U3284, and U3266 (including a repeat of U3266) were confirmed as true positives for Gram-negative pathogens (E. coli), and the GDS correctly identified these as Gram-negative. These positive samples were further subjected to antibiotic susceptibility testing (AST), as detailed in Table 3.
Table (3):
AST results (positive samples)
| Antibiotics |
Sample number |
|||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| U3274 |
U3284 | U3266 | ||||||||||
| POC | Disc diffusion | POC | Disc diffusion |
POC |
Disc diffusion |
|||||||
| O | I | Zone | I | 0 | I | Zone | I | O | I | Zone | I | |
| Piperacillin/Tazobactam | V | S | 23 | S | V | S | 27 | S | P | R | 20 | R |
| Ceftazidime | P | R | Absent | R | P | R | 20 | R | P | R | Absent | R |
| Cefepime | V | S | 14 | R | V | S | 20 | R | P | R | 13 | R |
| Aztreonam | P | R | 16 | R | P | R | 20 | R | P | R | 14 | R |
| Meropenem | V | S | 21 | S | V | S | 32 | S | V | S | 23 | S |
| Amikacin | V | S | 19 | S | V | S | 21 | S | V | S | 19 | S |
| Levofloxacin | V | S | 25 | S | V | S | 24 | S | V | S | 24 | S |
| Nitrofurantoin | V | S | 20 | S | V | S | 14 | S | P | R | Absent | R |
The five samples identified as contaminated by conventional methods (U3250, U3255, U3266, U3264, and U3298), the GDS produced false-positive results. This highlights a limitation of the GDS in distinguishing true infections from contaminated samples in certain cases (Table 2).
Urine samples contain Acinetobacter baumannii or yeast cells, the newly developed Gram Differentiation System (GDS) consistently produces a Pink/Pink combination. This occurs because the marker antibiotics used in this study are not recommended for these pathogens, as per CLSI guidelines. For example, sample U3266 (yeast cells) and sample U3298 (Acinetobacter baumannii) followed this pattern.
Gram categorization and antibiotic susceptibility testing (AST) results in urine samples and an Alamar Blue-based colorimetric system for point of-care (POC) detection (Table 2). The table compares colorimetric Gram categorization results with corresponding findings from AVMC laboratory culture-based identification.
The samples confirmed as true positives were further studied for AST detection. The Alamar Blue-based macro dilution sensitivity pattern showed an 87.5% similarity with disc diffusion results for samples U3274 and U3284 and a 100% similarity for sample U3266 (Table 3).
Antibiotic susceptibility testing (AST) results of positive urine samples using the point-of-care (POC) Alamar Blue-based colorimetric system compared with the conventional disc diffusion method. The Table 3 presents observation (O), interference (I), and zone diameter (mm) for selected antibiotics across three representative clinical samples. Note: O – Observation, I – Interference, V – violet, P – Pink, S – Sensitive, R – Resistant.
The limitations reported in previous literature are effectively addressed by the newly developed Alamar Blue assay, which incorporates both a Gram Differentiation System (GDS) and Antibiotic Susceptibility Testing (AST). For example, the UTI diagnostic kit developed by Sree Chitra Tirunal Institute for Medical Sciences and Technology and transferred to Agape Industries in 2024 demonstrated a major drawback it could detect only Gram-negative organisms and their corresponding AST profiles. This restricts its clinical utility, especially in primary healthcare settings where both Gram-positive and Gram-negative uropathogens are encountered.17 Similarly, another diagnostic kit developed in Chennai, within an animal containment zone, employed glycine-coated iron nanoparticles along with a methylene blue indicator. While this system used a combined panel of common antibiotics targeting both Gram-positive and Gram-negative organisms, it lacked pathogen-specific antibiotic guidance.18 According to Clinical and Laboratory Standards Institute (CLSI) guidelines, antibiotic susceptibility testing must be tailored to specific organism types.
The Alamar Blue assay system described in this study incorporates both a Gram differentiation system (GDS) and an antibiotic susceptibility testing (AST) component. For rapid and effective drug susceptibility determination at the point of care, identifying the Gram category of the infecting organism is often more critical than specific species-level identification. This is because antibiotic selection panels are primarily guided by whether the pathogen is Gram-positive or Gram-negative. Many previous studies have overlooked the inclusion of a GDS, particularly in veterinary applications, where broad-spectrum antibiotic panels are commonly used regardless of Gram classification. However, in the context of human clinical diagnostics especially when following Clinical and Laboratory Standards Institute (CLSI) guidelines the choice of antibiotics is more restrictive and must be precisely targeted. Without proper Gram categorization, drug susceptibility data may lead to inappropriate antibiotic selection. For example, prescribing a Gram-positive-targeted antibiotic for a Gram-negative infection can result in treatment failure due to poor efficacy or complete inactivity of the drug in the host environment.19
Initial evaluations with vancomycin (Gram-positive) and aztreonam (Gram-negative) demonstrated effective Gram categorization for Staphylococcus aureus, Enterococcus faecalis, and Pseudomonas aeruginosa. However, Escherichia coli posed challenges during broth microdilution assays. While agar testing confirmed expected resistance and sensitivity patterns, E. coli showed anomalous susceptibility to vancomycin and resistance to aztreonam in broth. To address this, antibiotic combinations were tested. Vancomycin plus linezolid was chosen for Gram-positive targeting, and aztreonam plus polymyxin B for Gram-negative organisms. The latter markedly improved E. coli classification, restoring its correct Gram-negative profile. Persistent inconsistencies with vancomycin/linezolid required further optimization. Concentrations below 8 µg/mL consistently yielded resistant profiles for E. coli, aligning with intrinsic resistance, whereas higher levels produced false sensitivity, likely from non-specific membrane disruption. The optimized system was validated across ATCC and clinical isolates, producing consistent, reliable colorimetric differentiation of Gram categories in accordance with reference standards.
During validation, the newly developed GDS demonstrated 100% accuracy in identifying the Gram category of all five tested ATCC strains when compared to conventional Gram staining. Additionally, the integrated AST module exhibited 97% accuracy, confirming its reliability in assessing drug susceptibility patterns. These findings highlight the robustness and compatibility of the developed system, supporting its potential for use in rapid, point-of-care diagnostics targeting urinary pathogens.
The newly developed Alamar Blue assay was further validated using real-world urine samples (n = 25). Among these, both the conventional diagnostic approach and the Alamar Blue assay identified 16 samples as negative for urinary tract infection (UTI), indicated by a consistent violet/violet color combination. Of the remaining nine samples, both methods confirmed four as true positives, with Escherichia coli identified as the etiological agent, reflected by the characteristic pink/violet color pattern. However, the remaining five samples were found to be improperly collected, exhibiting signs of contamination with mixed microbial flora. While the conventional method was able to identify these as contaminated by culture-based investigation, the Alamar Blue assay misclassified them as positive, resulting in five false-positive outcomes. This highlights a key limitation of the current assay system it lacks the ability to distinguish between true infections and contaminated specimens.
To address this limitation, it is crucial to emphasize the importance of proper urine sample collection. Advising patients to use a sample dispersion or midstream collection device can significantly reduce the risk of contamination.20 Additionally, integrating the Alamar Blue assay with a filtration-based diagnostic setup, as discussed earlier, may provide a more robust platform by enabling simultaneous detection of pyuria and bacteriuria.21 This system includes a dedicated sample container for the differential detection of pyuria and bacteriuria, along with a sealed plastic tray containing dedicated wells for Gram differentiation and antibiotic susceptibility testing. This refinement could improve diagnostic specificity and further enhance the assay’s applicability in point-of-care UTI detection systems. In contrast, the Alamar Blue assay presented in this study overcomes these limitations by enabling distinct antibiotic panels for Gram-positive and Gram-negative bacteria based on the integrated GDS system. This approach aligns with CLSI standards and provides more clinically relevant data. As a result, it offers a significant advantage in primary healthcare settings, empowering physicians to make more informed and accurate treatment decisions based on both rapid Gram categorization and tailored AST results.
The newly developed rapid concept for urinary tract infection (UTI) diagnosis was successfully evaluated using ATCC reference strains and 25 clinical urine samples. The antibiotic- and resazurin-based Gram differentiation system demonstrated 100% accuracy for five commonly encountered urinary pathogens. In addition, antibiotic susceptibility testing (AST) results showed strong agreement with the conventional Kirby–Bauer disk diffusion method. The present setup does not detect Acinetobacter baumannii, an occasionally encountered but clinically relevant urinary pathogen. The reliability of the new concept is highly dependent on proper urine sample collection, as inadequate collection may result in the identification of contaminant organisms rather than the true causative pathogen. A notable limitation of this Alamar Blue-based system is the potential for false-positive results arising from sample contamination. The observation of a pink–pink color pattern may indicate the presence of yeast, Acinetobacter species, or polymicrobial infection and should be interpreted with caution.
ACKNOWLEDGMENTS
The authors are thankful to the Vinayaka Mission’s Research Foundation (VMRF) and Aarupadai Veedu Medical College, Puducherry, for supporting this study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
PAA conceptualized the study. ACS, KK and EKR performed literature review and figure preparation. PAA performed the experiments and wrote the manuscript. LR, RS, and TS reviewed and revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
The study was approved by the Institutional Human Ethics Committee, Aarupadai Veedu Medical College and Hospital, Puducherry, India (Approval No: AV/IEC/2022/024).
- Mancuso G, Midiri A, Gerace E, Marra M, Zummo S, Biondo C. Urinary Tract Infections: The Current Scenario and Future Prospects. Pathogens. 2023;12(4):623.
Crossref - Aboobacker PA, Ragunathan L, Sanjeevi T, et al. Breaking boundaries in microbiology: customizable nanoparticles transforming microbial detection. Nanoscale. 2024;16(29):13802-13819.
Crossref - Geerlings SE. Clinical Presentations and Epidemiology of Urinary Tract Infections. Microbiol Spectr. 2016;4(5).
Crossref - Deltourbe L, Mariano LL, Hreha TN, Hunstad DA, Ingersoll MA. The impact of biological sex on diseases of the urinary tract. Mucosal Immunol. 2022;15(5):857-866.
Crossref - Yang X, Chen H, Zheng Y, Qu S, Wang H, Yi F. Disease burden and long-term trends of urinary tract infections: A worldwide report. Front Public Health. 2022;10.
Crossref - Salari N, Khoshbakht Y, Hemmati M, et al. Global prevalence of urinary tract infection in pregnant mothers: a systematic review and meta-analysis. Public Health. 2023;224:58-65.
Crossref - Mohapatra S, Panigrahy R, Tak V, et al. Prevalence and resistance pattern of uropathogens from community settings of different regions: an experience from India. Access Microbiol. 2022;4(2):000321.
Crossref - Hooton TM. Uncomplicated Urinary Tract Infection. N Engl J Med. 2012;366(11):1028-1037.
Crossref - Kline KA, Lewis AL. Gram-Positive Uropathogens, Polymicrobial Urinary Tract Infection, and the Emerging Microbiota of the Urinary Tract. Microbiol Spectr. 2016;4(2):10.
Crossref - Wilson ML, Gaido L. Laboratory Diagnosis of Urinary Tract Infections in Adult Patients. Clin Infect Dis. 2004;38(8):1150-1158.
Crossref - Endalamaw A, Erku D, Khatri RB, et al. Successes, weaknesses, and recommendations to strengthen primary health care: a scoping review. Arch Public Health. 2023;81(1):100.
Crossref - Gavina K, Franco LC, Khan H, Lavik JP, Relich RF. Molecular point-of-care devices for the diagnosis of infectious diseases in resource-limited settings – A review of the current landscape, technical challenges, and clinical impact. J Clin Virol. 2023;169:105613.
Crossref - Bermudez T, Schmitz JE, Boswell M, Humphries R. Novel technologies for the diagnosis of urinary tract infections. J Clin Microbiol. 2025;63(2):e0030624.
Crossref - Mambatta AK, Jayarajan J, Rashme VL, Harini S, Menon S, Kuppusamy J. Reliability of dipstick assay in predicting urinary tract infection. J Family Med Prim Care. 2015;4(2):265-268.
Crossref - Xu W, Venkat E, Koydemir CH. Optical point of care devices for diagnosis of urinary tract infections. Curr Opin Biomed Eng. 2023;28:100513.
Crossref - Zhang L, Wang B, Yin G, et al. Rapid Fluorescence Sensor Guided Detection of Urinary Tract Bacterial Infections. Int J Nanomed. 2022;17:3723-3733.
Crossref - Chakraborty D, Debnath F, Majumdar A, et al. Diagnostic validation study of rapid urinary tract infection diagnosis kit at peripheral health facilities of West Bengal, India. Sci Rep. 2024;14(1):297.
Crossref - Viswanathan K, Raj GD, Vadivoo VS, Kumanan K, Prabakaran R. Development of antibiotic selection kit towards veterinary applications using glycine passivated magnetic particles. Biosens Bioelectron. 2014;51:47-54.
Crossref - Apurba S Sastry SB. Essentials of Medical Microbiology. third. (Jangond AB, R D, eds.). Jaypee Brothers Medical Publishers. 2021. https://www.jaypeedigital.com/book/9789351529873
- Diviney J, Jaswon MS. Urine collection methods and dipstick testing in non-toilet-trained children. Pediatr Nephrol. 2021;36(7):1697-1708.
Crossref - Kunin CM, Buesching WJ. Novel Screening Method for Urine Cultures Using a Filter Paper Dilution System. J Clin Microbiol. 2000;38(3):1187-1190.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.