ISSN: 0973-7510
E-ISSN: 2581-690X
Tuberculosis (TB) is a serious and important public health issue to be addressed in India. Timely diagnosis of the drug resistance in tuberculosis is essential to ensure and initiate appropriate therapy. The detection of drug resistant Mycobacterium tuberculosis (MTB) in cases of smear-negative and clinical diagnosed pulmonary TB. A prospective case-control study was conducted on 473 pulmonary samples received at the tertiary care center from January 2019 to December 2019. All specimens were processed for microscopy and culture. CBNAAT- Gene Xpert and LPA Genotype MTBDRplus (VER 2.0) was used to confirm in smear-negative samples. Among the pulmonary samples, 52% smear-positive, and 48% smear-negative, 183 (43%) were found to be culture-positive by Lowenstein Jensen medium (LJ) and MGIT 960, 267 (63%) were positive CBNAAT and LPA n= 216 (51%) samples positive for the TUB band. The use of CBNAAT-Gene Xpert, Line Probe Assay Genotype MTBDR plus(VER 2.0) can be done from the samples directly and the diagnostic performance are more specific for detecting MTB in smear-negative specimens. This study suggests that LPA also helps in the diagnosis of MDR rapidly and in initiation of earlier treatment.
Smear-negative Samples, Line probe assay MTBDRplus, CBNAAT, Gene Xpert, MDR, Wild Type and Mutant
Tuberculosis (TB) a highly infectious disease.1 India is one (WHO, World Health Organization) among the high burdened countries responsible for more than 80%in the world prevalence of tuberculosis. Tuberculosis persists to be a one of the notifiable disease with high mortality and morbidity.2
The global incidence and the number of fatalities have slowly decreased over time, but are still very high in many parts of the world. Global TB epidemic is a major public health emergency. Every year ten million people develop the disease, and 1.4 million TB patients die (WHO, 2020).3
The increase of TB cases has threatened the current efforts to control TB with drug-resistant strains. Resistance to anti-tuberculosis medications emerges principally because of spontaneous mutations at unsurprising rates within the genome of M.tuberculosis.4 The MDR TB, resistance to isoniazid-INH and rifampicin-RIF are expanding and reported worldwide.5
Africa and Asia are the two regions with the largest burden of tuberculosis (TB) and affected for 95% of new TB cases, according to the World Health Organization (WHO, 2020).3,6 India has the highest number of TB patients in the world. India accounts for 27% of new TB cases developed globally, according to World Health Organization (WHO) estimates.7,8 WHO estimates that 4,84,000 cases of MDR/ rifampicin resistant TB were emerged, 6.2% of MDR-TB cases were identified as XDR-TB in 2018.7 Multidrug resistant TB has been defined as resistance to isoniazid and rifampicin with or without any other first line anti-TB drugs. About 3.4% of new cases and 18% of previously treated cases of TB in the world have MDR-TB or rifampicin-resistant TB. Additional resistance to fluoroquinolones and second-line injectables has been considered as extensively drug-resistant (XDR) TB.7
Although treatment in cases of drug resistant TB, MDR and XDR-TB is possible but expensive, lethal, longer than drug-susceptible TB, with higher rates of treatment disappointment and mortality.6 In MDRTB, genetic basis with mutation gene is responsible for resistance, at katG gene for isoniazid and rpoB genes for rifampicin.8
The rapid analysis of TB by a molecular strategy, the LPA was affirmed and supported by WHO to recognize the resistance to isoniazid and rifampicin.5 The limitations was overcome by the recent advanced form of LPA (The GenoType MTBDRplus LPA, version 2.0), a molecular diagnostic evaluation highly designed to recognize resistance especially to isoniazid, and rifampicin among the clinical suspected, smear-negative and directly from the sample.9
Gene Xpert not only detects M. tuberculosis but also confer mutational resistance to rifampicin with the help of three specific primers and ensure a high specificity. This cartridge-based testing platform with fully automated DNA detection used in TB diagnosis and to pick the rifampicin resistant within 2 hours. It’s faster in performance among the new generation of automated molecular diagnostics.10 This study compares the accuracy of CBNAAT, Gene Xpert and Line Probe Assays in the diagnosis of Tuberculosis in smear negative sputum samples.
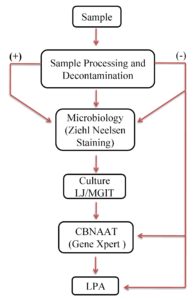
Pulmonary samples (473) were processed from January 2019 to December 2019 in a tertiary care center. The Institutional Human Ethical Committee approval and participant were enrolled after their written consent. The samples source were sputum 259 (61%), bronchial alveolar lavage 48(12%) followed by pleural fluid 76 (18%), and endotracheal aspirate 42(10%) and furthered processed. Figure 1 shows the steps followed for processing the samples. Nearly 48 pulmonary samples were excluded from the study either due to inadequate samples or contamination.
Figure 1. Sample processing steps
Note: Lowenstein Jensen medium- LJ, Line Probe Assay- LPA, Mycobacterium Growth Indicator Tube –MGIT, Drug Susceptibility Test -DST, Cartridge based nucleic acid amplification test–CBNAAT.
Sample Collection
In a wide-mouthed sterile container, sputum samples were collected (50-mL) (RNTCP -Revised National Tuberculosis Control Program guidelines).11 Microscopyby Ziehl-Neelsen (ZN) staining and decontamination were performed.12
Decontamination
Decontamination of all the specimens was done by the 5.0mL of NaCl solution (N-acetyl-L-cysteine), sodium hydroxide method (NaOH-NALC). After holding (15 min) at RT (Room Temperature), the specimens were neutralized with phosphate buffer saline (PBS) at pH 6 and placed in a cold centrifuged for 20 min at 10°C at 4500 rpm. Sterile phosphate buffer (1.5 ml) was used to suspend the pellets and collected for further analysis.13-15
Microscopic Examination
Zeihl-Neelsen Staining
The purulent part of the specimen was used to make a direct smear on a clean slide(2 × 1 cm). They were heat-fixed after air-drying and subjected for Zeihl-Neelsen staining (ZN stain) and graded according to RNTCP.12,16
Solid Medium: Lowenstein Jensen Medium (LJ)
After the decontamination samples, smear-positive and smear-negative samples were inoculated Lowenstein-Jensen Media. The processing of samples was done in Class II Bio-safety cabinet, BSL2. The slopes were incubated at 37°C after appropriate labeling with the date of inoculation. The slopes were assessed daily for growth. The time of appearance of first colony was noted, and slopes were additionally incubated for more growth.12
Liquid Medium: MycobacterialGrowth Indicator Tube (MGIT)
An MGIT-960 tube was inoculated with 0.8mL of PANTA and 0.5mL of sample and incubated at 37°C for 42 days (maximum) in MGIT-960 (Becton Dickinson) The final essential anti-TB medication concentrations, 0.1 g/ml INH and 1.0 g/ml RMP, were used during indirect DST. For MDRTB Line Probe Assay detection, specimens with smear positive, culture-positive, smear-negative sample were used.17,18
Cartridge Based Nuclear Acid Amplification Test (CBNAAT)
Sample (1 ml) was analyzed by CBNAAT on XpertMTB / RIF. There are 4 cartridges in the machine that was used for processing. The sampling reagent was added to the sample at 2:1 ratio, and kept at RT with intermittent shaking (15 min), according to standard operating procedure. 3 ml of sample transferred to the cartridge and inserted into the CBNAAT machine module.
The tests were then carried out on CBNAAT machines according to the user manual of the manufacturer. The automated analytics cartridge with results in 1 hour and 50 minutes. In the same setting, the detection of Mycobacteria and resistance to rifampicin was carried out. LPA was done for all samples to identify the rifampicin-resistance.19,20
GenoType MTBDRplus Ver2.0 Line-probe Assay
The GenoType MTBDRplus Ver 2.0 test was carried out as directed by the manufacturer instructions. Three stages make up the assay: For each of the three genes, the assay checks for the presence of wild type (WT) and mutant (MUT) probes followed by multiplex PCR (biotinylated primers) and reverse hybridization for RIF resistance, inhA for low-level INH resistance and katG for high-level INH resistance.13
Mycobacterial DNA was extracted and steps were as follows: 500 µl of the decontaminated material was centrifuged for 15 minutes at 10,000 rpm. The pellet was resuspended in 100 µl of sterile distilled water, warmed for 20 minutes at 95°C, sonicated for 15 minutes, then centrifuged for 8 minutes at 13, 000 rpm. The supernatant, which is the DNA solution, was put into another sterile 2 ml cryotube after centrifugation and kept at 20°C until ready for PCR. 21
Pre-made amplification mixes (amplification mix A and mix B) with all of the necessary components were used for PCR. 5 µl of MTB strain ATCC 27294 was used as a positive control, and 5 µl of sterile distilled water was utilized as a negative control. The sample DNA supernatant (5 µl) was utilized for Polymerase Chain Reaction in a 50 µl reaction mixture arranged by the kit instructions.9
Denaturation at 95°C for 15 minutes was followed by 10 cycles of initial denaturation at 95°C for 30 seconds and 58°C for 2 minutes, followed by 20 cycles of 95°C for 25 seconds, 53°C for 40 seconds, and 70°C for 40 seconds, and a final extension at 70°C for 8 minutes. The automated machine was used to hybridization (twincubator). Strips were removed after hybridization and washing, left to air dry, fixed on paper, and the findings were interpreted.20-22
The interpretation of the assay results was carried out, provided a template was used to align the strip containing 27 reaction zones (6 control probes and 21 mutation probes). Control by conjugate (CC), control by amplification (AC), M. tuberculosis complex (TUB), control of rpoB amplification, control of inhA amplification, and control of katG amplification are among the control probes. The presence of bands suggesting a mutation or the lack of at least one of the wild-type bands indicates that the sample is resistant to the antibiotic tested.23,24
A total of 473 pulmonary samples collected from the patient with clinical and radiological suspicion of tuberculosis. In 473 MTB suspected samples, nearly 48 pulmonary samples were excluded from the study either due to inadequate samples or contamination and under anti tuberculosis therapy (ATT). The sample sources are showed in Table 1.
Table (1):
Sample distribution in patients Smear positive, Culture positive and GenoType MTBDR-plus (VER 2.0).
| Source of Sample | No. of Sample n=425 (%) | Smear Positive ZN (%) | Culture Positive | CBNAAT Xpert MBT (%) | MTBDR-plus LPA (Positive Plus) | |
|---|---|---|---|---|---|---|
| LJ (%) | MGIT (%) | |||||
| Sputum | 259 (6) | 124 (48) | 67 (26) | 81 (31) | 113 (44) | 93 (36) |
| PF | 48 (12) | 29 (60) | 10 (21) | 13 (27) | 27 (56) | 19 (40) |
| BAL | 76 (18) | 38 (50) | 23 (30) | 27 (36) | 35 (46) | 32 (42) |
| ETA | 42 (10) | 29 (69) | 12 (29) | 19(45) | 28 (67) | 21 (50) |
| Total | 425 | 220 (52%) | 112 (26.3%) | 140 (33%) | 203 (48%) | 165 (39%) |
Note: PF – Pleural Fluid, BAL – Bronchial Alveolar Lavage, ETA – Endotracheal Aspirate, LJ – Lowenstein Jensen medium, CBNAAT-Cartridge based nucleic acid amplification test, MGIT- Mycobacterium Growth Indicator Tube, LPA- Line Probe Assay.
Nearly 289 and 136 samples were collected from male and female patients, respectively. Among 425 suspected cases, 346 samples received from the outpatient department and 79 had a history of contact with patients suffering from tuberculosis.
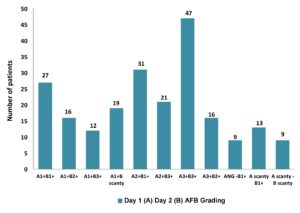
On microscopic examination, 220 (52%) were positive for AFB (ZN staining method), and 205 (48%) smear negative. Sample-based AFB grading is showed in (Figure 2). All samples were cultured (both LJ and MGIT). Out of 425 samples, 236 (56%) were found to be culture-positive and 189 (44%) negative.
In the study 425 sample, 220 (52%) were smeared positive (Table 1), and in them, 140 (64%) were culture positive. We found in study 205 (48%) smear negative samples to be processed for direct CBNAAT- Gene Xpert and LPA and in this smear-negative samples, 43 (21%) were culture positive (Table 2).
Table (2):
Performance of Genotype MTBDR plus VER 2.0 LPA for detection M. tuberculosis in smear negative sample.
| Source of Sample | Smear Positive | Culture Positive | CBNAAT Xpert MBT (%) MGIT (%) | MTBDR-plus LPA Positive (%) | |
|---|---|---|---|---|---|
| ZN (%) | LJ (%) | ||||
| Sputum | 124 (48) | 67 (26) | 81 (31) | 113 (44) | 93 (36) |
| PF | 29 (60) | 10 (21) | 13 (27) | 27 (56) | 19 (40) |
| BAL | 38 (7) | 7 (18) | 7 (18) | 14 (37) | 10 (26) |
| ETA | 13 (3) | 4 (31) | 6 (46) | 8 (62) | 8 (62) |
| Total | 205 | 34 (16.5%) | 43 (21%) | 64 (31.2%) | 51 (25%) |
Note: PF – Pleural Fluid, BAL – Bronchial Alveolar Lavage, ETA – Endotracheal Aspirate, LJ – Lowenstein Jensen medium, CBNAAT-Cartridge based nucleic acid amplification test, MGIT- Mycobacterium Growth Indicator Tube, LPA- Line Probe Assay.
Nearly 140 (64%) were MGIT 960 positive, In LPA, 165 (75%) were found with TUB band positive among the culture positive samples (Table 1) and 203 (93%) were CBNAAT positive. In smear-negative samples 43 (21%) positive for MGIT 960, 51 (25%) samples were found (TUB band) LPA positive, and 64 (31%) detected by CBNAAT.
MGIT 960 and CBNAAT (Rifampicin- RIF and Isoniazid- INH) sensitivity and resistant pattern were shown in (Table 3). In Nearly, 146(68%) were susceptible to both INH and RIF, 3 (1.8%) were MDR (resistant to both INH and RIF), 21 (13%) resistant to INH and 19 (12%) resistant to RIF (Table 3) in smear-positive and culture positive samples.
Table (3):
Drug susceptibility testing for Isoniazid and Rifampicin.
| Diagnostic Test | Smear Positive | Diagnostic Test | Smear Negative | ||||||
|---|---|---|---|---|---|---|---|---|---|
| RMP (S) | RMR (R) | INH (S) | INH (R) | RMP (S) | RMR (R) | INH (S) | INH (R) | ||
| MGIT n = 140 | 125 | 15 | 122 | 18 | MGIT n = 43 | 38 | 5 | 36 | 7 |
| X-pert n = 203 | 127 | 24 | – | – | X-pert n =64 | 46 | 18 | – | – |
| LPA n = 165 | 146 | 19 | 144 | 21 | LPA n = 216 | 43 | 8 | 42 | 9 |
Note: MGIT- Mycobacterium Growth Indicator Tube, DST – Drug Susceptibility Test, CBNAAT-Cartridge based nucleic acid amplification test, LPA- Line Probe Assay, INH: Isoniazid, RIF: Rifampicin.
In smear-negative, 18 (28%) were RIF resistant to CBNAAT (Figure 3), 43 (84%) susceptible to both INH and RIF, 4 (7.8%) MDR (resistant to INH and RIF), 9 (17%) resistant to INH, and 8 (16%) were resistant to RIF (Table 3). Interpretations of MTB resistant pattern by LPA are shown in Figure 4 and Table 4.
Table (4):
Interpretation of Mycobacterium tuberculosis Resistant pattern by Line Probe Assay (RMP and INH).
| No. Sample | RMP Resistance | INH Resistant | Result | ||||
|---|---|---|---|---|---|---|---|
| rpoB gene Missing | rpoB Mutant | KatG gene Missing | KatG Mutant | inhA gene missing | inhA Mutant | ||
| 1. | – | – | katGWT | – | – | – | Mono Resistant |
| 2. | – | – | katGWT | katGMUT 1 | – | – | Mono Resistant |
| 3. | – | – | – | – | inhAWT 2 | inhAMUT 1 | Mono Resistant |
| 4. | rpoBWT 7 | rpoBMUT 1 | – | – | – | – | Mono Resistant |
| 5. | rpoBWT 7 | – | – | – | inhAWT 2 | inhAMUT 1 | MDR Resistant |
| 6. | rpoBWT 7 | rpoBMUT 1 | – | – | – | – | Mono Resistant |
| 7. | rpoBWT 7 | katGWT | – | – | – | MDR Resistant | |
| 8. | – | – | – | – | inhAWT 2 | inhAMUT 1 | Mono Resistant |
| 9. | rpoBWT 7 | – | katGWT | katGMUT 1 | inhAWT 2 | – | MDR Resistant |
| 10. | – | – | katGWT | – | – | – | Mono Resistant |
| 11. | – | – | katGWT | – | – | – | Mono Resistant |
| 12. | – | – | – | – | inhAWT 2 | – | Mono Resistant |
| 13. | – | – | katGWT | – | – | – | Mono Resistant |
| 14. | rpoBWT 7 | rpoBMUT 1 | – | – | – | – | Mono Resistant |
| 15. | – | katGWT | katGMUT 1 | – | – | Mono Resistant | |
| 16. | – | – | – | – | – | inhAWT 2 | Mono Resistant |
| 17. | rpoBWT 7 | – | katGWT | katGMUT 1 | – | – | MDR Resistant |
| 18. | – | – | – | inhAWT 2 | – | Mono Resistant | |
| 19. | – | – | katGWT | – | – | – | Mono Resistant |
| 20. | rpoBWT 7 | – | – | – | inhAWT 2 | – | MDR Resistant |
| 21. | – | – | katGWT | – | – | – | Mono Resistant |
| 22. | rpoBWT 7 | – | – | – | – | – | Mono Resistant |
| 23. | – | katGWT | katGMUT 1 | – | – | Mono Resistant | |
| 16. | – | – | – | – | inhAWT 2 | inhAMUT 1 | Mono Resistant |
| 24. | rpoBWT | rpoBMUT 1 | katGWT | katGMUT 1 | – | – | MDR Resistant |
| 25. | rpoBMUT 1 | – | – | – | Mono Resistant | ||
| 26. | rpoBWT | – | katGWT | – | – | – | MDR Resistant |
| 27. | – | – | – | – | inhAWT 2 | – | Mono Resistant |
Note: INH: Isoniazid, RIF: Rifampicin, MDR: Multi Drug Resistant, MUT: Mutant, WT: Wild Type.
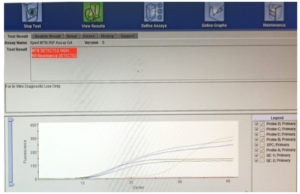
Figure 3. CBNAAT on Xpert-MTB/RIF.
Note: The graph lines showed the Rifampicin resistant from the smear negative sample.
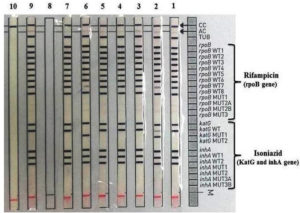
Figure 4. Band patterns of GenoType MTBDR-plus VER 2.0 line probe assay.
Positive MTB complex presence of ‘TUB’ band in all strips
Line 1: MTB susceptible to both RIF and INH
line 2: MDRTB with RIF/INH resistant (absence of wild-type band rpoBWT7, KatGWT and presence of a mutation band KatG MUT1.
lines 3-5: MTB susceptible to both RIF and INH
line 6: MDRTB with RIF/INH resistant (absence of wild-type band rpoBWT7, KatGWT and with a mutation band KatG MUT1
Line 7: MTB susceptible to both RIF and INH
line 9: Positive control Mycobacterium tuberculosis strain, ATCC 27294
line 10: Negative control (sterile distilled water). MDRTB: Multidrug resistant tuberculosis,
INH: Isoniazid, RIF: Rifampicin, MTB: Mycobacterium tuberculosis.
In this study, a total of 425 samples; the male population (68%) was more predominant than female (32%), with an average of 18-65 years. New cases were 81% in them 19% with a history of contact. Our study was in concurrence with Abyot Meaza et al.23 Out of these 182, 118 (65%) patients were male, and 64 (35%) were female patients. The median age 40 in a range of (20- 65 years) was not similar to the study of Kandha kumara et al.25 who has reported the median age of the patient was 26 years (range 1 year to 68 years). Among 425 samples 220 (52%) showed smear positive for AFB , 205 (48%) were negative.
All negative samples were inoculated on LJ media and MGIT culture, 183 (43%) showed culture positive. This is similar to the study of Namratha W et al., and documented that among, 265 showed AFB positive, and 115 showed AFB negative.26 In drug susceptibility testing, 7% isoniazid (INH) and 6.3% rifampicin (RIF) resistance which was similar to Krairerk Suthum et al.18 Of 183 (43%) culture positive, 20 RIF-resistant strains (by MGIT 960 array) and in them 25 (14%) isoniazid-resistant strains were identified similar to Chao-Ju Chen et al.27
In 425 pulmonary samples (Sputum, Pleural fluid Bronchial alveolar lavage, Endotracheal aspirate samples – CBNAAT Gene Xpert), 158 samples were negative. This study found the rifampicin resistance rate 28%. CBNAAT could identify 64 cases (31.2%) that smeared negative was similar to Dash Manoranjan et al.20 In our study out of the cohort of 205 samples smear-negative patients, MTB was detected in 64 patients by CBNAAT, and 18 RIF-resistant, and 46 were RIF sensitive. CBNAAT could identify 64 cases (31.2%) from smear negative samples was similar to Dash Manoranjan et al.20 The CBNAAT showed rifampicin sensitivity pattern in concurrent to Vishal Chopra et al.28
Sputum smear was AFB positive in 124 (48%) patients, and CBNAAT detected MTB in 113 (44%) subjects. Among them, 12 had rifampicin resistance similar to Anushree Chakrabortyet al.29 Out of 6.3% were resistant by about were rifamcin resistant LPA, whereas 9.85 were rifampicin-resistant by CBNAAT. A total of 209/425 (49%) showed negative for MTB by LPA and 158/425 (37%) showed negative for MTB in CBNAAT which was not in concurrence with Tripathi et al.19
All AFB negative specimens were inoculated on LJ Media, 34 (17%) showed culture positive. Pleural fluid of 48(12%) sample were subjected for TB culture (both LJ & MGIT), 13 (27%) were culture positive, and 8 (42%) were negative, 27 (56%) were positive by LPA. Similar results documented by Muhammad Irfan et al.30
In our study in 216 (51%) Genotype MTBDRplus LPA (v2.0),detected 7 (3.2%) MDR, 27 (13%) resistant to RIF and 30 (14%) resistant to INH, 189 (88%) were susceptible to both INH and RIF. The above documentation of drug-resistant TB was similar to the study of Barnard et al.,(86.3% susceptible to both RIF and INH).28 Our study is in concurrence to susceptibility (95%) and mono resistant (4.2%) to RIF, (91%) susceptibility and (9%) resistant to INH, 7 (2.4%) MDR strains with Stephen et al.24 In the present study, 9 (3%) cases with RIF-WT7 probe was detected and similar to Ritu Singhal et al.21 In INH resistant, 6(2.1%) showed katG gene mutations in the inhA gene promoter region detected by LPA and this resistance pattern was not in concurrence with Sara Siddiqui et al.33
In our study, 51/216 (24%) of the smear-negative sample showed TUB band (LPA). Our study similar to Binit Kumar Singh et al.34 The use of Genotype MTBDRplus LPA (v2.0) directly from the smear-negative samples and susceptibility to RIF (84%), INH(82%) and detection of mono resistant strains to RIF (16%), to INH (18%) and MDR 3 (6%) were similar to Syed Beenish Rufai et al.35 This study supports the use of Genotype MTBDRplus LPA (v2.0) for detection of newer cases and MDR from the directly from pulmonary sample.
CBNAAT can be essential in sputum negative patients as a diagnostic modality, for early diagnosis and medication. It is less time consuming, but it can detect only rifampicin resistance. Genotype MTBDRplus VER 2.0 is a rapid diagnostic method for TB detection in smear-negative specimens and promotes accurate diagnosis of both RMP and INH resistance, susceptibility to drugs within a short turnaround time (4 hours). It provides rapid information to opt for early treatment and contain disease spread.
ACKNOWLEDGMENTS
The authors would like to thank Prof. Dr. R. Murugesan (Advisor-Research) Karpaga Vinayaga Institute of Medical Sciences and Research Center for critical review of the manuscript and valuable suggestions. The authors would also like to thank Chettinad Academy of Research and Education (CARE), Chettinad Hospital and Research Institute (CHRI), Chennai, India for providing the research fellowship.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This research was supported by Chettinad Academy of Research and Education (CARE), Chennai, India.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the Institutional Human Ethical Committee (141a/IHEC/06-18), Chettinad Academy of Research and Education, Chennai, India.
- Dean T. Jamison, Hellen Gelband, Susan Horton, Prabhat Jha, Ramanan Laxminarayan Charles N. Mock, Major Infectious Diseases, Disease Control Priorities, Third Edition. 2017;6.
Crossref - Use of high burden country lists for,TB by WHO in the post-2015 era, WHO/HTM/ TB/2015.29
- Rutaihwa LK, Zwyer M, Borrell S, et al, Local adaptation in populations of Mycobacterium tuberculosis endemic to the Indian Ocean Rim. F1000 Research 2021;10:60.
Crossref - Rai DK, Pandey S. A Hospital-Based Cross-sectional Study on Clinico demographic Characteristic of Extrapulmonary Tuberculosis Cases Coming to a Tertiary Hospital of Bihar. Indian J Community Med. 2018;2020.
- Seung KJ, Keshavjee S, Rich ML. Multidrug-Resistant Tuberculosis and Extensively Drug-Resistant Tuberculosis, Cold Spring Harb Perspect Med. 2015;5(9):a017863.
Crossref - Companion Handbook to the WHO Guidelines for the Programmatic Management of Drug-Resistant Tuberculosis; 2014,ISBN: 978924154880 9, WHO/HTM/TB/2014.11.
- Singh AV, Singh S, Yadav A, et al. Genetic variability in multidrug-resistant Mycobacterium tuberculosis isolates from patients with pulmonary tuberculosis in North India. BMC Microbiol. 2021;21(1):123.
Crossref - Isakova J, Sovkhozova N, Vinnikov D, et al. Mutations of rpoB, katG, inhA and ahp genes in rifampicin and isoniazid-resistant Mycobacterium tuberculosis in Kyrgyz Republic. BMC Microbiol. 2018;18(1):22.
Crossref - Shah A, Rodrigues C. The Expanding Canvas of Rapid Molecular Tests in Detection of Tuberculosis and Drug Resistance. Astrocyte. 2017;4:34-44.
Crossref - Sachdeva K, Shrivastava T. CBNAAT: A Boon for Early Diagnosis of Tuberculosis-Head and Neck. Indian J Otolaryngol Head Neck Surg. 2018;70(4):572-577.
Crossref - Yadav RN, Singh BK, Sharma SK, et al. Comparative Evaluation of GenoType MTBDRplus Line Probe Assay with Solid Culture Method in Early Diagnosis of Multidrug Resistant Tuberculosis (MDR-TB) at a Tertiary Care Centre in India. PLoS ONE. 2013;8(9):e72036.
Crossref - Stinson KW, Eisenach K, Matsumoto M, et al. Mycobacteriology Laboratory Manual, First Edition. Global Laboratory Initiative. 2014.
- Revised National Tuberculosis Control Programme Laboratory, Guidelines for Quality Assurance of smear microscopy for diagnosing tuberculosis. 2005.
- Kazemian H, K-Y J, Bahador A, et al. Efficacy Of Line Probe Assay In Detection Of Drug-Resistant Pulmonary Tuberculosis In Comparison With GeneXpert And Phenotypic Methods In Iran And Genetic Analysis Of Isolates By MIRU-VNTR. Infect Drug Resist. 2019;12:3585-3593.
Crossref - Schwartz D, Shafran I, Romero C, et al. Use of short-term culture for identification of Mycobacterium avium subsp. paratuberculosis in tissue from Crohn’s disease patients. Clin Microbiol Infect. 2000;6(6)303-307.
Crossref - Vijdea R, Stegger M, Sosnovskaja A, Andersen AB, Thomsen VO, Bang D. Multidrug resistant tuberculosis: rapid detection of resistance to rifampin and high or low levels of isoniazid in clinical specimens and isolates. Eur J Clin Microbiol Infect Dis. 2008;27(11):1079-1086.
Crossref - Djoudi F, Benremila D, Kassa S, et al. Diagnosis and Drug Susceptibility of Mycobacterium tuberculosis from Pulmonary Specimens at Pasteur Institute of Algeria: Comparative Study between Classic Lowenstein-Jensen Culture and BACTEC MGIT 960 System. EC Pulmonology and Respiratory Medicine. 2019;8.4.
- SuthumK, Samosornsuk W, Samosornsuk S. Characterization of katG, inhA, rpoB and pncA in Mycobacterium tuberculosis isolates from MDR-TB risk patients in Thailand. J Infect Dev Ctries. 2020;14(3):268-276.
Crossref - Tripathi R, Sinha P, Kumari R, Chaubey P, Pandey A, Anupurba S. Detection of rifampicin resistance in tuberculosis by molecular methods: A report from Eastern Uttar Pradesh, India. Indian J Med Microbiol. 2016;34(1):92-94.
Crossref - Dash M, Samal S, Swain TR, Behera B, Sahu S, Mahapatra S. Diagnostic yield of cartridge based nucleic acid amplification test in Mycobacterium tuberculosis in a tertiary care medical college and hospital of Southern Odisha, India. Int J Res Med Sci. 2019;7(12):4460-4463.
Crossref - Singhal R, Anthwal D, Kumar G, et al. Genotypic characterization of ‘inferred’ rifampin mutations in GenoType MTBDRplus assay and its association with phenotypic susceptibility testing of Mycobacterium tuberculosis. Diagn Microbiol Infect Dis. 2020;96(4):114995.
Crossref - Kumar P, Balooni V, Sharma BK, Kapil V, Sachdeva KS, Singh S. High degree of multi-drug resistance and hetero-resistance in pulmonary TB patients from Punjab state of India. Tuberculosis. 2014;94(1):73-80.
Crossref - Meaza A, Kebede A, Yaregal Z, et al. Evaluation of genotype MTBDRplus VER 2.0 line probe assay for the detection of MDR-TB in smear positive and negative sputum samples. BMC Infect Dis. 2017;17(1):280.
Crossref - Stephen S, Muzhizhizhi D, Dhibi N, et al. Validation of the GenoType MTBDRplus Ver 2.0 Assay for Detection of Rifampicin and Isoniazid Resistance in Mycobacterium tuberculosis Complex Isolates at UZCHS CTRC TB Research Laboratory. Int J Mycobacteriol. 2019;8(1):83-88.
Crossref - Gandhi K, Selvaraj S, Kannaiyan K. Isolation of Diverse Mycobacterium Tuberculosis Strains Employing Automated and Conventional Culture from Lymphadenitis in a Tertiary Care Center, Pondicherry, India. Journal of Research in Medical and Dental Science. 2019;7(3):164-168.
- Nandihal NW, Anand MK. Molecular Detection of Rifampicin and Isoniazid Resistance and Characterization of Mutations in Mycobacterium tuberculosis Complex using Line Probe Assay. Int J Curr Microbiol App Sci. 2019;8(1):465-471.
Crossref - Chen C-J, Yang Y-C, Huang H-H, Chang TC, Lu P-L. Evaluation of a membrane hybridization array for detection of Mycobacterium tuberculosis complex and resistance to isoniazid and rifampin in sputum specimens, mycobacterial liquid cultures, and clinical isolates. Kaohsiung J Med Sci. 2019;35(10):615-623.
Crossref - Barnard M, van Pittius NCG, van Helden PD, Bosman M, Coetzee G, Warren RM. The diagnostic performance of Genotype MTBDRplus Version 2 line probe assay is equivalent to the XpertMTB/RIF assay. J Clin Microbiol. 2012;50(11):3712-3716.
Crossref - Chakraborty A, Ramaswamy S, Shivananjiah AJ, Puttaswamy RB, Chikkavenkatappa N. The role of genexpert in the diagnosis of tubercular pleural effusion in India. Adv Respir Med. 2019;87(5):276-280.
Crossref - Irfan M, Idrees F, Jabeen K, Zubairi ABS, Butt S, Hasan R. Accuracy of genotype MTBDRplus line probe assay in patients with tuberculous pleural effusion: comparison with clinical and culture based diagnosis. Infect Dis. 2020;52(4):235-241.
Crossref - SiddiquiI S, Brooks MB, Malik AA, et al. Evaluation of GenoType MTBDRplus for the detection of drug-resistant Mycobacterium tuberculosison isolates from Karachi, Pakistan. 2019;14(8):e0221485.
Crossref - Singh BK, Sharma SK, Sharma R, et al. Diagnostic utility of a line probe assay for multidrug resistant-TB in smear-negative pulmonary tuberculosis. PLoS ONE. 2017;12(8):e0182988.
Crossref - Rufai SB, Kumar P, Singh A, Prajapati S, Balooni V, Singh S. Comparison of Xpert MTB/RIF with Line Probe Assay for Detection of Rifampin-Mono resistant Mycobacterium tuberculosis. J Clin Microbiol. 2014;52(6):1846-1852.
Crossref
© The Author(s) 2022. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.