ISSN: 0973-7510
E-ISSN: 2581-690X
Brucellosis, a reported zoonotic disease caused by several species of the genus Brucella, is endemic to many countries, including Morocco, and poses a major public health challenge. Available studies and national surveillance data highlight its persistent presence and impact, with an incidence rate that is difficult to estimate owing to under-reporting of cases. Human diagnosis is mainly based on serological tests (Rose Bengal and ELISA) without the routine implementation of molecular techniques, which are faster and more specific. To our knowledge, in Morocco, no study has demonstrated the use of polymerase chain reaction (PCR) as a routine diagnostic tool. This study aimed to develop, test, and optimise a multiplex real-time PCR assay capable of detecting all Brucella spp. using the conserved bcsp31 gene to differentiate between B. melitensis and B. abortus. Using the KoMa plasmid as an internal extraction control, this study assessed the analytical sensitivity of the assay, which showed a detection limit of ≤10 plasmid copies and 5-10 genomic copies per reaction, including its specificity, which reached 100%, with no amplification observed for non-Brucella strains. This pilot study evaluated the diagnostic performance of the assay using 30 suspected brucellosis samples. All 17 were confirmed to be positive by ELISA and Rose Bengal tests, whereas 13 were negative. Seventeen positive samples were confirmed by real-time PCR, which showed 100% agreement (95% confidence interval [CI]: 98%). The results showed that all positive human cases studied were caused by B. melitensis. This fast and reliable method is a promising tool for monitoring and managing the rapid clinical diagnosis of human brucellosis. However, validation on a larger scale is required.
Brucella spp., B. melitensis, B. abortus, Multiplex PCR, Real-Time, Molecular Diagnostics, Morocco
Brucellosis is a globally important zoonotic disease caused by various species of the genus Brucella.1 The two main species responsible for human brucellosis are Brucella abortus and Brucella melitensis.2 This disease imposes a heavy socioeconomic burden on many countries.3 These bacteria are transmitted to humans through consumption of contaminated products or occupational exposure.4 This pathology remains endemic in many countries, including Morocco.5
Human brucellosis is frequently underreported because of its non-specific clinical presentation, which ranges from mild symptoms to severe complications, including its variable incubation period and the general lack of awareness regarding the disease.6-8 Diagnosis cannot be based solely on clinical criteria.7 Consequently, diagnostic delays are a major challenge, often allowing the infection to progress from an acute to a chronic condition, thereby increasing the risk of complications.9 Therefore, a rapid and accurate laboratory confirmation is essential.
Isolation of Brucella spp. remains the gold standard for diagnosis. However, the culture sensitivity is often low, potentially leading to false-negative results.10 Furthermore, isolation requires Biosafety Level 3 (BSL-3) facilities and highly skilled personnel owing to the significant risk of laboratory-acquired infections.3 Alternatively, serological tests detecting lipopolysaccharides (LPS) are widely implemented.11 However, their specificity is limited by cross-reactivity with other Gram-negative bacteria that share similar antigens, such as Yersinia enterocolitica.7
To overcome these limitations, molecular methods, particularly qualitative and quantitative polymerase chain reactions (PCR), have been developed to improve diagnostic capabilities.12-14 Multiplex real-time PCR offers rapid screening and allows for the simultaneous detection of the genus Brucella and differentiation between B. melitensis and B. abortus.15,16 These methods offer significant advantages in terms of speed and cost-effectiveness compared with traditional screening.7,15
In Morocco, accurate incidence estimates are hampered by under-reporting.17 Diagnosis relies primarily on serological methods despite their limitations, whereas molecular approaches are rarely used in routine protocols.18 To our knowledge, the routine use of PCR for brucellosis diagnosis has not yet been established in Morocco.
This study’s primary objective was to develop and evaluate molecular tools for diagnosing Brucella infection. Specifically, this study optimised a multiplex real-time PCR targeting the bcsp31 sequence for genus-level detection and differentiation of two major species involved in human infection: B. melitensis and B. abortus. A synthetic plasmid (KoMa) was used as an internal extraction control (IEC) to validate the analytical procedure.
The second objective of this pilot study was to validate the analytical performance of the assay, specifically its specificity and limit of detection (LOD), and evaluate its utility in clinical samples from cases confirmed by ELISA and Rose Bengal assays.
Sources of strains and DNA
A panel of Brucella and non-Brucella strains was selected from the American Type Culture Collection (ATCC) Collection and maintained at the Laboratory of Medical Microbiology of Epidemic Diseases, National Institute of Hygiene (Table 1). The genomic DNA of B. melitensis and B. abortus, used as positive controls, was provided by the Robert Koch Institute (RKI), Berlin.
Table (1):
List of bacterial strains used to evaluate assay specificity
Bacterial Strains |
Origin (Reference) |
No. of Strains |
|---|---|---|
Escherichia coli |
ATCC 25922 |
3 |
Haemophilus influenzae |
ATCC 49247 |
2 |
Neisseria meningitidis |
ATCC 13090 |
2 |
Pseudomonas aeruginosa |
ATCC 27853 |
2 |
Staphylococcus aureus |
TCC 25923 |
2 |
Staphylococcus epidermidis |
ATCC 12228 |
2 |
Streptococcus pyogenes |
ATCC 19615 |
2 |
Streptococcus pneumoniae |
ATCC 49619 |
2 |
Enterococcus faecalis |
ATCC 51299 |
2 |
Listeria monocytogenes |
ATCC 7644 |
2 |
Shigella sonnei |
ATCC 29930 |
2 |
Yersinia enterocolitica |
ATCC 27729 |
3 |
Klebsiella pneumoniae |
ATCC 700603 |
2 |
Salmonella enteritidis |
ATCC 13076 |
2 |
Bacillus subtilis |
ATCC 6633 |
2 |
Candida albicans |
ATCC 14053 |
2 |
DNA B. melitensis |
ATCC 23456 |
2 |
DNA B. abortus S19 |
NCBI ID 430066 |
2 |
*ATCC: American Type Culture Collection *NCBI: National Centre for Biotechnology Information
Evaluation of analytical specificity
Analytical specificity was evaluated using a panel of Brucella and non-Brucella strains described in Table 1. All the strains were cultured under standard laboratory conditions. Genomic DNA was extracted using a commercial nucleic acid purification kit according to the manufacturer’s instructions.
The multiplex real-time PCR assay employed specific primers and hydrolysis (TaqMan) probes targeting Brucella spp., with an internal amplification control (IAC) included in each reaction to monitor PCR inhibition. The specificity was defined as the percentage of non-Brucella strains that yielded negative results (without amplification). This was calculated using the following formula:
Analytical Specificity (%) = [TN / (TN + FP)] × 100
Where TN represents the number of True Negatives (non-Brucella strains with no amplification), and FP represents the number of False Positives (non-Brucella strains showing amplification).
Analytical sensitivity assessment
Analytical sensitivity was determined using serial 10-fold dilutions of Brucella melitensis genomic DNA, ranging from an initial concentration of 1 ng/µL down to 1 pg/µL. Each dilution was tested in duplicate under the optimised multiplex amplification conditions. A standard calibration curve was generated by plotting the recorded threshold cycle (Ct) values against the logarithm of the initial DNA concentration. The LOD was defined as the lowest DNA concentration consistently detected with a reliability of ≥95% among replicates.
Clinical samples
In the present study, 30 human serum samples were selected from the Biobank of the Medical Laboratory of Epidemic Diseases, National Institute of Hygiene (Rabat, Morocco). The samples were obtained from patients with suspected brucellosis in southern Morocco. Before this study, all samples were characterised using Rose Bengal and ELISA; 17 were confirmed to be positive for brucellosis, and 13 were confirmed to be negative.
Genomic DNA extraction from bacteria and serum
DNA was extracted from 200 µL of serum or bacterial culture using the DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Five microlitres of a plasmid containing the artificial KoMa2 sequence were added to each sample as an IEC.19 DNA was eluted in 100 µL of elution buffer. DNA concentration and purity were assessed using a NanoDrop One spectrophotometer (Thermo Fisher Scientific, USA). Extracts were stored at -20 °C until analysis.
Real-time PCR assays
A multiplex TaqMan real-time PCR assay was developed to simultaneously target the bcsp31 gene (specific to the genus Brucella) and the species-specific sequences of B. abortus and B. melitensis. A separate singleplex assay targeting the KoMa2 sequence was used to monitor the IEC. Primers and probes were designed by the RKI (Berlin, Germany) and synthesised by Qiagen (Alameda, USA); the sequences and concentrations are detailed in Table 2.
Table (2):
Primers and oligonucleotide probes used in multiplex real-time PCR for the detection of Brucella spp., B. abortus, and B. melitensis with KoMa2 extraction control19,20
| Target | Primer/Probe | Sequence (5′-3′) |
|---|---|---|
| Brucella spp. | Brucella spp.–F | GCT CGG TTG CCA ATA TCA ATG C |
| Brucella spp.–R | GGG TAA AGC GTC GCC AGA AG | |
| Brucella spp.–TM | FAM-AAA TCT TCC ACC TTG CCC TTG CCA TCA-BHQ-1 | |
| B. abortus | B. abortus-F | GCG GCT TTT CTA TCA CGG TAT TC |
| B. abortus-R | CAT GCG CTA TCA CGG TAT TC | |
| B. abortus-TM | HEX-CGC TCA TGC TCG CCA GAC TTC AAT G-BHQ-1 | |
| B. melitensis | B. melitensis-F | AAC AAG CGG CAC CCC TAA AA |
| B. melitensis-R | CAT GCG CTA TGA TCT GGT TAC | |
| B. melitensis-TM | Cy5-CAG GAG TGT TTC GGC TCA GAA TAA TCC ACA-BHQ-2 | |
| KoMa (IEC) | KoMa-F | GGTGATGCCGCATTATTACTAGG |
| KoMa-R | GGTATTAGCAGTCGCAGGCTT | |
| KoMa-TM | HEX-TTCTTGCTTGAGGATCTGTCGTGGATCG-BHQ-2 |
F: Forward primer; R: Reverse primer; TM: TaqMan Probe
PCR reactions were prepared in a final volume of 25 µL, containing 6.25 µL of TaqMan™ Environmental Master Mix (Applied Biosystems), 0.75 µL of each primer, 0.25 µL of each hydrolysis probe, 8.50 µL of nuclease-free water, and 5 µL of template DNA. All samples were assayed in duplicates. Sterile PCR-grade water and purified Brucella genomic DNA were used as the negative and positive controls, respectively.
Thermal cycling was performed on a QuantStudio 5 Real-Time PCR System (Applied Biosystems) with the following protocol: initial activation at 95 °C for 10 min, followed by 40 cycles of denaturation at 95 °C for 15 sec and annealing/extension at 60 °C for 60 sec. Data acquisition and Ct calculation were performed using QuantStudio Design & Analysis Software.
Internal quality control
Each PCR run included negative (nuclease-free water) and positive controls. An IAC was incorporated into each sample reaction. Plasmid standards were also included to allow for semi-quantitative evaluation and to serve as additional positive controls.
Contamination testing
To assess potential cross-contamination, 6 controls (3 positive and 3 negative) were processed simultaneously during DNA extraction and amplification. These controls were arranged in an alternating pattern on a PCR plate (positive/negative) to monitor the material transfer between wells.
Practicability
The robustness, reproducibility, and clinical performance of the multiplex assay were evaluated through an interlaboratory comparison organised by the RKI. All participating laboratories used the same protocol for Brucella genus identification and species differentiation (B. abortus and B. melitensis). The concordance between centres was used to validate the assay’s practicability across different laboratory conditions.
Statistical analysis
Quantitative data (fluorescence signals and Ct values) were generated using the QuantStudio 5 software (Applied Biosystems). Diagnostic accuracy parameters, including sensitivity, specificity, and overall concordance with reference serological assays, were calculated using Microsoft Excel. The results are reported with 95% Confidence Intervals (95% CIs). Cohen’s kappa coefficient was calculated to quantify the agreement between the multiplex real-time PCR assay and the serological methods for the 30 clinical samples evaluated.
Interpretation criteria
The results were interpreted as follows: a sample was considered positive if an amplification curve was detected in the Brucella spp. channel with a Cycle Threshold (Ct) <38. A sample was considered negative if no signal was detected in the Brucella channel, provided that the IEC (KoMa) had a Ct < 35, confirming valid extraction and amplification. Samples showing no signal for either target were classified as invalid and were retested.
Clinical performance and detection
Among the 30 suspected cases tested in the present cohort, 17 tested positive by multiplex real-time PCR, demonstrating 100% clinical sensitivity and concordance with serological results. All 17 positive samples were identified as B. melitensis (Table 3). B. abortus DNA was not detected in any of the clinical samples.
Table (3):
Multiplex real-time PCR amplification results for Brucella controls, non-Brucella strains, and clinical serum samples
Sample type |
Brucella spp. (Target) |
B. melitensis (Target) |
B. abortus (Target) |
|---|---|---|---|
B. melitensis (Control) |
+ |
+ |
– |
B. abortus (Control) |
+ |
– |
+ |
Non-Brucella strains |
– |
– |
– |
Serum samples (n = 30) |
17 (Positive) |
17 (Positive) |
0 (Positive) |
Note: ‘+’: amplification detected; ‘-’: no amplification
The KoMa internal control was successfully detected in all samples (mean Ct: 35 ± 3). The Ct values for the Brucella-specific targets ranged from 25-33, indicating moderate to low bacterial loads. The Ct values for B. melitensis specific targets ranged from 30-38. No amplification was observed in the 13 seronegative samples or the healthy controls, confirming 100% clinical specificity. In the studied population, the genus-specific multiplex real-time PCR assay showed perfect agreement with ELISA results, with a Cohen’s kappa coefficient of 1.00, indicating excellent concordance (Table 4).
Table (4):
Agreement between genus-specific multiplex real-time PCR and ELISA for human sera (n = 30)
Method |
Cohen’s κ |
95% Confidence Interval |
P-value |
|---|---|---|---|
Genus-specific PCR vs ELISA |
1.00 |
1.00a-1.00 |
Notes: κ = Cohen’s kappa coefficient
Statistical Agreement: The genus-specific multiplex real-time PCR demonstrated perfect agreement with ELISA results (κ = 1.00; P < 0.001) in the studied population
Analytical specificity
No cross-reactivity was observed. All Brucella strains were correctly detected, whereas none of the non-Brucella strains generated specific signals. Consequently, analytical specificity was calculated to be 100%, as shown in Table 3.
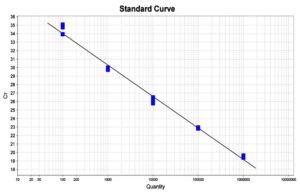
Standard curve and LOD
The assay demonstrated high linearity with a correlation coefficient (R²) of 0.99 and an amplification efficiency of 95%. The LOD was 5-10 genome copies per reaction for B. melitensis genomic DNA and 10 copies per reaction for plasmid standards (Figure).
Figure. Standard curve to correlate the DNA concentration of B. melitensis in copies/µL with the cycle threshold (Ct)
Sample size and concordance
Multiplex real-time PCR successfully detected Brucella DNA in all seropositive samples, demonstrating 100% concordance with serological results. These findings are summarised in Table 5.
Table (5):
Multiplex real-time PCR detection rates in seropositive serum samples (n = 17)
Sample Group |
Test 1 (Positive) |
Test 2 (Positive) |
Total (Positive) (n) |
Total Tested (N) |
Detection Rate |
|---|---|---|---|---|---|
Seropositive Sera |
17 |
17 |
17 |
17 |
100% |
Note: The assay demonstrated 100% agreement with reference serological results across technical replicates
The analytical performance of the multiplex real-time PCR assay has proven to be robust for the routine diagnosis of Brucella spp., which is consistent with the findings reported in several studies.21 Under the conditions of this study, a specificity of 100% excluded cross-reactions with related bacterial species and a LOD of approximately 5-10 genomic copies per reaction allowed for the identification of very low DNA loads, thereby supporting early patient management.
The concordance between multiplex real-time PCR and serological testing is encouraging. However, the rather wide 95% confidence interval (90%-100%) reflects the restricted sample size and indicates high uncertainty around this estimate. Nevertheless, the low detection limit and high specificity underscore the assay’s utility as a fast and accurate diagnostic tool.21 The multiplex real-time PCR developed here demonstrated analytical sensitivity comparable to that of the most reliable methods reported in the literature, and absolute specificity against non-Brucella pathogens.12 Furthermore, inclusion of the KoMa plasmid as an internal control ensured the validity of the DNA extraction and amplification steps, thereby minimising false-negative results.22
Human brucellosis is characterised by broad and non-specific clinical manifestations. The clinical presentation alone does not allow for a definitive diagnosis.23,24 Untimely treatment also increases the risk of chronic complications. Consequently, early and accurate diagnosis is crucial for the treatment, control, and eradication of the disease and serves as the cornerstone of prevention and surveillance in humans.25 Fast, reliable, sensitive, specific, and automated detection systems for Brucella spp. are urgently needed to facilitate early diagnosis and adequate antibiotic therapy.2,26
Considering the limitations of the conventional diagnostic approaches for human brucellosis.27 In this context, the multiplex real-time PCR assay evaluated in this study provides a rapid, sensitive, and highly specific alternative for detecting and differentiating Brucella species.28 The assay demonstrated excellent clinical performance, full concordance with serological findings, and reliable species-level identification, highlighting its potential utility as a complementary diagnostic tool to support timely diagnosis and improve brucellosis surveillance in endemic regions such as Morocco, where diagnosis currently relies primarily on serological methods.29,30
Owing to these constraints, multiplex real-time PCR has emerged as an integrated diagnostic solution that combines speed, safety, and accuracy. Unlike conventional methods, this technique does not require the handling of live agents, thereby reducing the biosafety requirements of BSL-2 for DNA extraction.31 The optimised protocol provides results in <3 hrs, compared to several days for culture, with an analytical sensitivity of 5-10 genomic copies/reaction.32,33 The simultaneous detection of specific targets (bcsp31 for Brucella spp., omp2a for B. melitensis, and alkB for B. abortus) ensured 100% specificity, as demonstrated in panels that included 20 non-Brucella strains.21,34
Recent studies increasingly support the added diagnostic value of molecular approaches for brucellosis.3 Several studies have highlighted the utility of PCR-based methods as confirmatory tools, particularly for complex clinical presentations or when serological results are inconclusive.35 In endemic settings, molecular assays are recognised as valuable adjuncts to conventional diagnostics, providing improved specificity and aiding in the interpretation of ambiguous serological findings.36
Regarding species distribution, B. melitensis was identified in 100% of positive cases in this pilot cohort. These observations are consistent with data reported in various publications, indicating that B. melitensis is responsible for the majority of human cases.37,38 Recent publications have noted that routine identification of Brucella species involved is often lacking.39 Currently, data on Brucella species circulating in various hosts in Morocco are limited, which hinders the effective study and management of the disease.36 Identifying the specific species involved in human infections is essential for facilitating epidemiological traceability and applying necessary health measures.2
The application of this assay to diverse samples (blood, fixed tissues, and joint fluids) extends its utility to focal forms of the disease that are often culture-negative.40 In summary, the adoption of multiplex real-time PCR meets modern molecular diagnostic requirements, including speed, safety, and analytical accuracy. Its future integration as a complementary diagnostic approach, consistent with internationally recognised guidelines, could support surveillance and control strategies in endemic regions, such as Morocco.41
External quality assessments (EQA) are essential for clinical laboratories to ensure reliable and accurate test results.42,43 The multiplex real-time PCR test was evaluated through inter-laboratory comparisons, achieving a score of 100%. This guaranteed the effectiveness and sensitivity of the assay for the detection and differentiation of B. melitensis and B. abortus, as recommended by the World Health Organisation (WHO) for detecting Brucella DNA.44
Based on the available literature, this study is among the first to use multiplex real-time PCR for the species-specific diagnosis of human brucellosis (B. melitensis/B. abortus) in a clinical context in Morocco. The results confirmed that all human cases studied were caused by B. melitensis. The assay demonstrated high sensitivity and specificity, allowing for rapid screening and accurate differentiation. However, the limited number of samples prevented extrapolation to national epidemiological trends. The multiplex real-time PCR evaluated in this pilot study is promising for addressing the constraints of conventional methods. However, further multicentre validation involving a larger cohort of clinically suspected cases across diverse geographic regions and varying temporal frames is warranted before its integration into the national diagnostic algorithm. Future investigations should include additional clinically relevant sample matrices to further confirm the validity of this test.
To our knowledge, this is the first report on the molecular diagnosis of human brucellosis caused by Brucella melitensis and B. abortus in Morocco. The results confirmed that all positive human cases in this cohort were attributable to B. melitensis. The developed multiplex real-time PCR assay demonstrated high sensitivity and specificity, offering a reliable complementary tool for the rapid screening and accurate differentiation of the two primary species involved in human infection. The integration of this technique into diagnostic and epidemiological surveillance protocols, particularly within livestock control programs, is recommended as part of an integrated “One Health” approach. Further studies are warranted to improve the understanding of the distribution of Brucella species among human and animal populations in Morocco, and to strengthen national control strategies for this persistent zoonosis.
ACKNOWLEDGMENTS
The authors express their sincere gratitude to the Robert Koch Institute (RKI, Germany) for their valuable technical support and responsiveness throughout this study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
AQ, NA, BB, and RC conceptualized the study. AQ, NA, FO, KH, JH, and RC applied methodology. AQ, NA, BB, and RC performed validation. AQ, NA, BB, and RC wrote, reviewed and revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
This study was funded by the German Biosafety Program.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- World Health Organization, Food and Agriculture Organization, eds. Brucellosis in Humans and Animals. Geneva, Switzerland: World Health Organization; 2006. https://www.who.int/publications/i/item/9789241547130
- Ghssein G, Ezzeddine Z, Tokajian S, et al. Brucellosis: Bacteriology, pathogenesis, epidemiology and role of the metallophores in virulence: a review. Front Cell Infect Microbiol. 2025;15:1621230.
Crossref - Franc KA, Krecek RC, Häsler BN, Arenas-Gamboa AM. Brucellosis remains a neglected disease in the developing world: a call for interdisciplinary action. BMC Public Health. 2018;18(1):125.
Crossref - CDC. Brucellosis Reference Guide: Exposures, Testing and Prevention. Natl Cent Emerg Zoonotic Infect Dis Dis. 2017:40. www.cdc.gov/brucellosis/pdf/brucellosi-reference-guide.pdf. Accessed May 27, 2021.
- Kazemi S, Afshar S, Karami M, et al. Association between risk of brucellosis and genetic variations in MicroRNA-146a. BMC Infect Dis. 2021;21(1):1070.
Crossref - Qasmaoui A, Belkadi B, Ohmani F, Halout K, Charof R, Hamamouchi J. New approach needed for diagnosis of human brucellosis in Morocco. E3S Web Conf. 2021;319:01018.
Crossref - Becker GN, Tuon FF. Comparative study of IS711 and bcsp31-based polymerase chain reaction (PCR) for the diagnosis of human brucellosis in whole blood and serum samples. J Microbiol Methods. 2021;183:106182.
Crossref - Zhai J, Peng R, Wang Y, et al. Factors Associated With Diagnostic Delays in Human Brucellosis in Tongliao City, Inner Mongolia Autonomous Region, China. Front Public Health. 2021;9:648054.
Crossref - Pang C, Liu L, Liu T, Zhang N, Zhao L. Delayed diagnosis of pediatric brucellosis: a case study and literature review. Annals of Medicine & Surgery. 2025;87(6):3880-3884
Crossref - Thakur S, Bedi JS, Singh R, Gill JPS, Arora AK, Kashyap N. Quantitative polymerase chain reaction based quantification of Brucella DNA in serum of pre- and post-therapeutic occupationally exposed infected human population. J Infect Public Health. 2018;11(4):514-520.
Crossref - Gumaa MM, Osman HM, Omer MM, El Sanousi EME, Godfroid J, Ahmed AM. Seroprevalence of brucellosis in sheep and isolation of Brucella abortus biovar 6 in Kassala State, Eastern Sudan. Rev Sci Tech-Off Int Epizoot. 2014;33(3):957-965.
Crossref - Yu WL, Nielsen K. Review of Detection of Brucella sp. by Polymerase Chain Reaction. Croat Med J. 2010;51(4):306-313.
Crossref - Moussa IM, Omnia ME, Amin AS, Ashgan MH, Selim SA. Evaluation of the currently used polymerase chain reaction assays for molecular detection of Brucella species. Afr J Microbiol Res. 2011;5(12):1511-1520.
Crossref - Freire ML, Machado De Assis TS, Silva SN, Cota G. Diagnosis of human brucellosis: Systematic review and meta-analysis. PLoS Negl Trop Dis. 2024;18(3):e0012030.
Crossref - Lopez-Goni I, Garcia-Yoldi D, Marin CM, et al. Evaluation of a Multiplex PCR Assay (Bruce-ladder) for Molecular Typing of All Brucella Species, Including the Vaccine Strains. J Clin Microbiol. 2008;46(10):3484-3487.
Crossref - Probert WS, Schrader KN, Khuong NY, Bystrom SL, Graves MH. Real-Time Multiplex PCR Assay for Detection of Brucella spp., B. abortus, and B. melitensis. J Clin Microbiol. 2004;42(3):1290-1293.
Crossref - Nawana TB, Ezzine H, Cherkaoui I, et al. Brucellosis at the animal-human-environment interface in Morocco, 2002-2019: a descriptive analysis. Pan Afr Med J One Health. 2021;6:13.
Crossref - Faddane K, Moumni H, Cherkaoui I, et al. Seroprevalence of human brucellosis in Morocco and associated risk factors. Vet World. 2022;15(9):2224-2233.
Crossref - Kirchner S, Kramer KM, Schulze M, et al. Pentaplexed Quantitative Real-Time PCR Assay for the Simultaneous Detection and Quantification of Botulinum Neurotoxin Producing Clostridia in Food and Clinical Samples. Appl Environ Microbiol. 2010;76(13):4387-4395.
Crossref - Kang SI, Her M, Kim JW, et al. Advanced Multiplex PCR Assay for Differentiation of Brucella Species. Appl Environ Microbiol. 2011;77(18):6726-6728.
Crossref - Kattar MM, Zalloua PA, Araj GF, et al. Development and evaluation of real-time polymerase chain reaction assays on whole blood and paraffin-embedded tissues for rapid diagnosis of human brucellosis. Diagn Microbiol Infect Dis. 2007;59(1):23-32.
Crossref - Schrader C, Schielke A, Ellerbroek L, Johne R. PCR inhibitors – occurrence, properties and removal. J Appl Microbiol. 2012;113(5):1014-1026.
Crossref - Khoshnood S, Pakzad R, Koupaei M, et al. Prevalence, diagnosis, and manifestations of brucellosis: A systematic review and meta-analysis. Front Vet Sci. 2022;9:976215.
Crossref - Bai Q, Li H, Wu X, Shao J, Sun M, Yin D. Comparative analysis of the main outer membrane proteins of Brucella in the diagnosis of brucellosis. Biochem Biophys Res Commun. 2021;560:126-131.
Crossref - Aly Reda SEH, El-Messiah MAA, Madkour LA, Kassem AEM. Serology versus Real Time PCR in the Diagnosis of Human Brucellosis. J Pure Appl Microbiol. 2017;11(2):677-683.
Crossref - Che LH, Qi C, Bao W, et al. Monitoring the course of Brucella infection with qPCR-based detection. Int J Infect Dis. 2019;89:66-71.
Crossref - Sayar MS, Bulut D, Haykir Solay A. Two Cases of Laboratory Acquired Brucellosis. Mediterr J Infect Microbes Antimicrob. 2018;7:32.
Crossref - Qureshi KA, Parvez A, Fahmy NA, et al. Brucellosis: epidemiology, pathogenesis, diagnosis and treatment–a comprehensive review. Ann Med. 2023;55(2):2295398.
Crossref - Qasmaoui A, Natoubi S, Bougharouine M, et al. Serological diagnosis of human brucellosis in Morocco and prospects for advanced diagnostic techniques. Iran J Microbiol. 2025;17(5):695-701.
Crossref - Li J, Jin J, Liu Y, et al. Development and evaluation of multiple droplet digital PCR method for specific detection and differentiation of Brucella melitensis and Brucella abortus. Animals. 2026;16(4):566.
Crossref - Elbehiry A, Aldubaib M, Al Rugaie O, et al. IS711 sequencing of Brucella melitensis and Brucella abortus strains, and use of microchip-based real-time PCR for rapid monitoring. J King Saud Univ – Sci. 2023;35(2):102468.
Crossref - Mitka S, Anetakis C, Souliou E, Diza E, Kansouzidou A. Evaluation of Different PCR Assays for Early Detection of Acute and Relapsing Brucellosis in Humans in Comparison with Conventional Methods. J Clin Microbiol. 2007;45(4):1211-1218.
Crossref - Zamanian M, Jahani E, Mahmoudi H. Multiplex PCR Assay for the Simultaneous Detection of the Brucella Genus in Human Whole Blood and Serum. Open Microbiol J. 2020;14(1):242-246.
Crossref - Liu H, Jiang H. Advances in the application of molecular diagnostic techniques to brucellosis. Biomedical and Environmental Sciences, 2024;37(7):790-794.
Crossref - Nazir S, Farooq M, Khan R, et al. Comparative evaluation of diagnostic tests for brucellosis in humans and animals: a meta-analytical approach. Vet Sci. 2025;12(7):638.
Crossref - Dadar M, Alamian S, Tadayon K, Ashford RT, Whatmore AM. Molecular characterization of zoonotic Brucella species isolated from animal and human samples in Iran. Acta Trop. 2022;229:106363.
Crossref - Lounes N, Cherfa MA, Le Carrou G, et al. Human Brucellosis in Maghreb: Existence of a Lineage Related to Socio-Historical Connections with Europe. PLoS ONE. 2014;9(12):e115319.
Crossref - Sahin H, Erdenlig Gurbilek S, Ozcan N, Saytekin AM, Yucetepe AG. Molecular typing of Brucella species in human brucellosis cases from Eastern Turkiye. Revista Científica de la Facultad de Ciencias Veterinarias. 2025;XXXIV(3):1-5.
Crossref - Ksibi B, Smaoui F, Ben Ayed N, et al. Genomic analysis of Brucella melitensis isolates recovered from humans in south Tunisia over 35 years between 1988 and 2022. BMC Microbiol. 2025;25(1):98.
Crossref - Loubet P, Magnan C, Salipante F, et al. Diagnosis of brucellosis: Combining tests to improve performance. PLoS Negl Trop Dis. 2024;18(9):e0012442.
Crossref - Jiang H, Feng L, Lu J. Updated Guidelines for the Diagnosis of Human Brucellosis — China, 2019. China CDC Wkly. 2020;2(26):487-489.
Crossref - Laudus N, Nijs L, Nauwelaers I, Dequeker E. The Significance of External Quality Assessment Schemes for Molecular Testing in Clinical Laboratories. Cancers. 2022;14(15):3686.
Crossref - Manjengwa J, Zakaryan A, Orelle A, et al. Development and implementation of National External Quality Assurance Programs in a One Health approach: The Armenian experience. One Health. 2021;13:100351.
Crossref - World Health Organization. WHO Manual for Organizing a National External Quality Assessment Programme for Health Laboratories and Other Testing Sites. World Health Organization; 2016. https://iris.who.int/handle/10665/250117. Accessed June 28, 2024.
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.