ISSN: 0973-7510
E-ISSN: 2581-690X
Antimicrobial-resistant bacterial disease has been recognized by WHO as a critical global health threat, Staphylococcus aureus posing particular concern due to its role in severe septic infections and bacteremia. This concern has led to an urgent need for alternative therapies, such as nanotechnology and plant-based approaches. The study focused on synthesizing plant-based silver nanoparticles from aqueous extract of Catharanthus roseus stem (CrS-AgNPs) prepared via the decoction process. The formation of CrS-AgNPs was confirmed by UV-Vis spectroscopy with a stable peak at 426 nm, followed by further characterization using FTIR and XRD analyses. The antimicrobial potential against pathogenic S. aureus was evaluated using the resazurin assay. Furthermore, SDS-PAGE was performed for whole cell protein profiling, and RT-PCR was used to analyze the expression of the virulence gene phospholipase C (plc). The results showed six functional peaks in the FTIR spectrum, while XRD analysis revealed a prominent peak at 38.33°, indicating a face-centered cubic geometry, and the calculated particle size is 26.74 nm. MIC and MBC were determined as 0.04 µg/mL and 0.32 µg/mL, respectively. While SDS-PAGE indicated protein-level alterations, RT-PCR showed no significant change in plc gene expression. Silver nanoparticles synthesized from C. roseus stem were successfully obtained, exhibiting functional groups of C≡N, H-C=O, and O-H, which contribute to their antimicrobial properties. Further investigation is needed to understand the underlying mechanisms and explore their therapeutic potential.
Catharanthus roseus, UV-Vis Spectroscopy, Pathogenic Staphylococcus aureus, qPCR, plc gene
Antimicrobial resistance is responsible for about 1.27 million deaths globally (Global Burden of Disease, 2019), and the World Health Organization also has concerns about antimicrobial-resistant Staphylococcus aureus (S. aureus), which is recognized as a leading cause of mortality and morbidity worldwide. Over the past two decades, antimicrobial-resistant S. aureus strains have predominantly developed due to the extensive and uncontrolled use of antibiotics.1,2 Pathogenic S. aureus, a coagulase- and gram-positive bacterium, can colonize various body sites, including the nasal passages and soft tissues, and is frequently associated with bacteremia and sepsis.3,4 However, S. aureus exhibits an extensive repertoire of virulence factors, such as surface-anchored proteins, which enable it to evade the host immune response.5,6 Virulence-associated regulatory genes, including agr, SaeRS, and vraRS, enhance its ability to invade and survive within the host.7,8 A pivotal virulence factor of S. aureus is phospholipase C (plc),9 whereas the agr quorum-sensing system and the SrrAB two-component regulatory system are critical regulators of their gene expression.10 Although broad-spectrum antibiotics, including methicillin, vancomycin, and ciprofloxacin are commonly used to cure infections caused by S. aureus.11
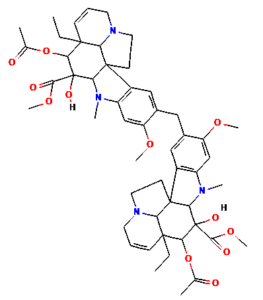
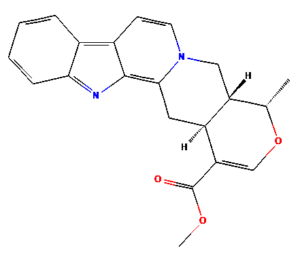
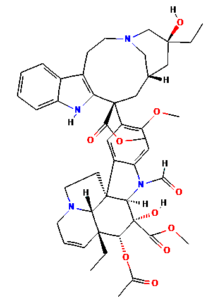
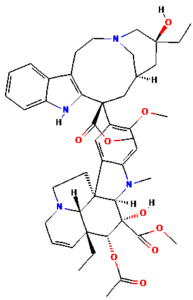
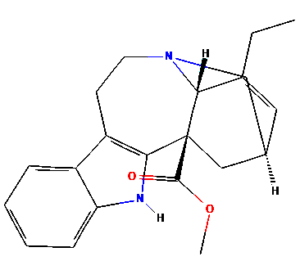
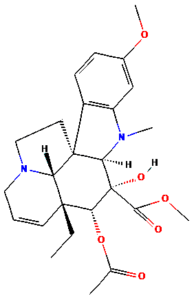
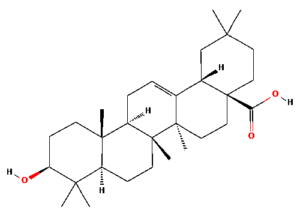
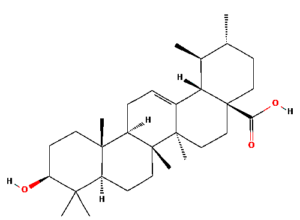
The ongoing challenge of combating bacterial infection is associated with diminished antibiotic efficacy and limited alternative therapy strategies. The molecular mechanisms underlying bacterial virulence and resistance pathways offer valuable insights into the development of innovative antimicrobial strategies.12 Ethnomedicinal plants have promising avenues for novel therapeutics, among which Catharanthus roseus (C. roseus) of the Apocynaceae family, traditionally used in Ayurveda, is rich in bioactive secondary metabolites such as alkaloids, flavonoids, and terpenoids. C. roseus exhibits a wide range of pharmacological properties, such as antibacterial, anti-inflammatory, anticancer, and anti-diabetic properties.13 Numerous alkaloids, including vindoline, catharanthine, serpentine, and vindolicine, have been identified, while vinca alkaloids such as vinblastine and vincristine are extensively used in cancer treatment. Additionally, phytoconstituents, such as oleanolic acid and ursolic acid, have demonstrated antimicrobial properties (Table 1).14-16
Table (1):
Phytoconstituents extracted from Catharanthus roseus with their 2D-chemical structures
Plant-based nanoformulations have led to attention because of their potential for antimicrobial properties and their ability to offer safer, non-toxic alternatives for both human and environmental health. Metallic nanoparticles of silver have gained significant interest due to their distinctive properties, such as high electrical conductivity and surface area, and ease of use.22 The green synthesis of plant-based silver nanoparticles (AgNPs) offers an alternative method to overcome the physical and any toxic chemical methods and is eco-friendly, safe, and cost-effective. The current exploration of AgNPs emphasizes their significant role in addressing antimicrobial resistance while identifying key challenges and future prospects for their application in biomedicine.23-26
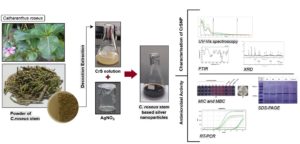
In this study, AgNPs were synthesized from an aqueous C. roseus stem (CrS) extract via the decoction method (Figure 1). Ultraviolet-visible (UV-Vis) spectroscopy confirmed AgNPs formation, while Fourier Transform Infrared Spectroscopy (FTIR) was used for characterization of functional groups, and X-ray Diffraction (XRD) was used for morphology and crystalline structure. Antibacterial efficacy against pathogenic S. aureus was evaluated. Whole-cell protein profiling was conducted via sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), followed by plc gene expression analysis to assess the impact of AgNPs.
Identification and extraction of C. roseus stem by decoction method
C. roseus (sadabahar/periwinkle), a widely available medicinal plant in Asia, was collected from the Sehore-Bhopal region of Madhya Pradesh, India, and authenticated with reference no. 2/19424R on 24/04/2024 at Vedanta Testing & Research Laboratory, Bhopal. The dried material was crushed and made into powder and stored in an airtight, dark glass container at room temperature. An aqueous CrS extract was prepared by boiling 50 g of stem powder in 200 mL of distilled deionized water at 80 °C for 5 min using the decoction method. To enhance phytoconstituents from the extraction method, the mixture was mixed and stirred at 200 rpm for 5 hours, then filtered to obtain the extract. The resulting yellowish CrS extract was used for AgNPs synthesis.
Silver nanoparticles synthesis via CrS extract
A 1 mM AgNO₃ solution (>99.9%, Sigma Aldrich) was prepared in 45 mL of distilled water, mixed with 5 mL of CrS extract, and adjusted to pH 7. The mixture was stirred and heated at 85-90 °C for 2 hours in a heat plate condenser assembly. The capping process was monitored every 30 min using UV-Vis spectroscopy (Genesys 180, Thermo Scientific) to assess the reduction of silver (Ag⁺) ions by the CrS extract. The solution gradually changed from bright yellow to black, indicating the formation of CrS extract-based AgNPs, and the sample was cooled to room temperature. The sample was centrifuged at 9000 rpm for 30 min at 25 °C, washed with deionized water to remove impurities, and then dried at 80 °C for 6 hours.27,28 The dried CrS-AgNPs were carefully collected and stored in a sterile glass container for further analysis.
Characterization of CrS-AgNPs using FTIR and XRD
The synthesized CrS-AgNPs were characterized using FTIR and XRD. The FTIR (IRAffinity-1S, Shimadzu) analysis was done in the wavenumber range of 4000-400 cm-1, and samples were ground into KBr pellets.29 The crystalline structure of the synthesized CrS-AgNPs was determined by the XRD method using a diffractometer (MINIFLEX, RIGAKU-JAPAN) equipped with copper Kα radiation (λ = 1.5406 Å), scanned in a 2θ range of 3°-80° at a rate of 5°/min. The instrument operated at 40 kV and 20 mA, using copper Kα radiation. The crystallite size of the NPs was estimated using Debye-Scherrer’s equation:30
D = Kα / lcosθ
where D is the mean crystallite size (nm), K is the shape constant (0.9), λ is the X-ray wavelength (0.15406 nm), β is the full width at half maximum (FWHM) of the diffraction peak in radians, and θ is the Bragg angle in radians.31
Minimum Inhibitory Concentration (MIC) and Minimum Bactericidal Concentration (MBC) determination against S. aureus
The MIC assay assessed the antibacterial activity of CrS-AgNPs against pathogenic S. aureus (MTCC96), procured from the MTCC and Gene Bank, Chandigarh, India. Bacterial culture was revived and maintained at 37 °C for 24 hours according to standard protocol. The culture was grown in Mueller-Hinton broth (MH) and standardized to 5 × 105 CFU/mL using the McFarland standard. CrS-AgNPs (3 mg/mL in DMSO) were tested in a 96-well microtiter plate. The CrS-AgNPs solution was serially diluted in bacterial culture across the wells. Ciprofloxacin (30 µL, 1 mg/mL) was used as the standard drug, while the untreated bacterial culture and the blank served as the positive and negative controls, respectively. Culture plates were incubated for 16-18 hours at 37 °C. After that, resazurin (0.015%, 30 µL) was added and incubated for 3 hours; a color change from blue to pink indicated bacterial growth. MBC was determined by treated samples that were streaked onto MH agar plates and incubated at 37 °C for 18 hours. Tests were done in triplicate.
Whole-cell protein profiling via SDS-PAGE
Changes in the whole-cell protein profile of S. aureus were evaluated using SDS-PAGE by comparing CrS-AgNPs-treated and untreated cells. Band patterns indicated changes in protein expression. Two groups were used: one treated with CrS-AgNPs at the MIC concentration and one untreated control. Samples were incubated for 2, 4, 8, and 12 hours. Whole-cell proteins were extracted using the alkaline lysis method followed by Laemmli SDS buffer. Cell lysates were incubated with loading dye (1:1) at 95 °C for 10 min and stored at -20 °C. SDS-PAGE analysis was run at 120 V using a 12% resolving gel and 5% stacking gel prepared with acrylamide. Each well was loaded with 10 µL of protein extract along with a prestained protein ladder (MBT092-10LN, HiMedia, India). After gels were stained with Coomassie Brilliant Blue and destained using a solution of 10% methanol and acetic acid until clear protein bands became visible.32
qPCR of plc gene expression
The plc gene, encoding thermonuclease and associated with virulence in S. aureus, is documented under ENA accession number SUJ92450.1. The mRNA expression of plc was analyzed by qPCR in CrS-AgNPs-treated and untreated S. aureus cells. Primers for amplification of plc virulence gene were designed to generate a 284 bp amplicon using the forward primer 5′-TGGGCAGATAACGCGACATT-3′ and reverse primer 5′-ATCAGCCAACCCGTTCTAG-3′. The 16S rRNA gene (UHAE01000001.1), as the housekeeping control, produced a 204 bp amplicon with the forward primer 5′-GAGTACGACCGCAAGGTTGA-3′ and reverse primer 5′-CCCAACATCTCACGACACGA-3′.33 Cell concentration was adjusted to 1×108 CFU/mL using the McFarland standard and treated with CrS-AgNPs (0.04 µg/mL). The culture was incubated in a rotary shaker at 37 °C for 12 hours. Total RNA was extracted using an RNA extraction kit (HiMedia, India) following the manufacturer’s instructions, and its purity was measured spectrophotometrically using the A260/A280 ratio (Genesys 180, Thermo Scientific). Complementary DNA (cDNA) was synthesized from the extracted RNA (HiMedia, India). The RNA and random hexamers were heated at 65 °C for 5 min and then rapidly cooled as per protocol. The reaction was processed in a thermal cycler at 25 °C for 5 min, 42 °C for 60 min, and 70 °C for 5 min, followed by a final hold at 4 °C. The resulting cDNA was evaluated using RT-PCR. Amplification of 5 µg cDNA was performed using the HiScript Two-Step RT-PCR Kit (HiMedia, India), following the manufacturer’s instructions. SYBR Green (SYBR, ROX, SMOBiO) was used as the fluorescent indicator for the polymerase reaction. For the amplification of 16S rRNA and plc gene, the thermal cycling conditions included 94 °C for 1 min, 54 °C for 15 sec, and 72 °C for 45 sec, followed by a final extension at 72 °C for 4 min over 30 cycles. Relative expression was calculated using the 2ΔΔCt method.
Statistical analysis
Statistical data were analyzed in Excel 2016 using one-way ANOVA (mean ± standard deviation, P < 0.05).
Spectroscopic analysis
A UV-Vis spectrophotometer was used to analyze the optical behavior of synthesized AgNPs. CrS-AgNPs formation was monitored by recording surface plasmon resonance (SPR) absorbance between the wavelengths of 200-800 nm at 30 min intervals. Previous studies reported that AgNPs display maximum absorption in the UV-Vis range of 350-500 nm.34 In the present study, Figure 2 displays the absorption band of the green-synthesised CrS-AgNPs. A color change appeared after 60 min, with the reaction stabilising at 150 min, marked by a characteristic dark brown color. The UV-Vis spectrum of CrS-AgNPs showed a prominent SPR peak near 426 nm, confirming nanoparticle formation. The observed SPR characteristics were influenced by factors such as particle size, shape, composition, and morphology. The initial absorbance at 426 nm of the mixture was 0.27, which gradually increased to 0.37, 0.39, and 0.68. A maximum absorbance of 0.73 was observed at 150 min, confirming the complete reduction of silver nitrate. According to the IMPPAT database, C. roseus extract contains phytoconstituents such as alkaloids, flavonoids, terpenoids, and phenols that aid in the capping of AgNPs.35 Its reported antibacterial, anticancer, antiseptic, and anti-inflammatory properties make C. roseus based AgNPs a promising candidate for drug development. pH adjustment was crucial for the formation of CrS-AgNPs, as confirmed by UV-Vis spectra showing complete reduction of silver nitrate within 150 min.
Figure 2. (a) UV-visible absorption spectrum demonstrating a characteristic peak at 426 nm, confirming the biosynthesis of silver nanoparticles using Catharanthus roseus stem (CrS-AgNPs) extract. (b) Photograph of the synthesized CrS-AgNPs in their dried powdered form
XRD analysis
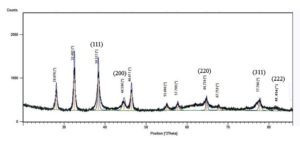
XRD is used to determine the crystalline lattice structure or amorphous nature of the biosynthesized silver nanoparticles. The XRD pattern of CrS-AgNPs exhibited distinct peaks at 2θ values of 28.08°, 32.48°, 38.33° (111), 44.59° (200), 46.41°, 55.05°, 57.71°, 64.74° (220), 67.76°, 77.75° (311), and 82.85° (222). These were analyzed using the ICSD database (File No. 064996/Ref. No. 01-087-0719) as a reference to correlate the 2θ angle values with 5 characteristic intensities (Table 2 and Figure 3). Previous studies have reported that the highest intensity peak at 38.33° corresponds to the (111) plane, clearly indicating the formation of AgNPs.36 The obtained diffraction pattern confirmed that the CrS-AgNPs possessed a face-centered cubic crystal structure, resulting from the reduction of silver ions by the aqueous CrS extract. Additional peaks observed in the XRD pattern may be attributed to biological components present in the extract. Using the Debye-Scherrer equation, the average crystallite size was calculated to be 26.74 nm, based on a 2θ angle at 38.33° with a FWHM of 0.315°. The measurements were performed within a 2θ range of 20°-100° using copper Kλ radiation (λ = 1.5406 Å).
Table (2):
Summary of XRD results for biosynthesized Catharanthus roseus stem-based silver nanoparticles
Sample |
2θ peak angle |
hkl, Diffraction plane |
Peak width (β, radians) |
Cosθ |
Crystalline size (D, nm) |
|---|---|---|---|---|---|
CrS-AgNPs |
38.33 |
111 |
0.32 |
0.95 |
26.74 |
Figure 3. XRD patterns display insights into the crystalline nature and structural information of silver nanoparticles synthesized using the aqueous extract of the Catharanthus roseus stem
FTIR analysis of CrS-AgNPs and CrS extract
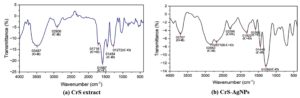
Functional group identification and detection of vibrational shifts in AgNPs derived from CrS extract were determined using FTIR spectroscopy. As shown in Figure 4, the FTIR spectra exhibited notable shifts between the CrS extract and CrS-AgNPs, confirming the active participation of the extract phytoconstituents in nanoparticle formation. A similar kind of shift was reported in a previous study.37 The FTIR spectrum of CrS-AgNPs showed multiple peaks, indicating the presence of various biomolecular functional groups. A strong peak at 2296 cm-1 confirmed C≡N stretching (nitrile groups), an indication of NP formation,38 while 2677 cm-1 corresponded to H-C=O stretching (aldehyde/formyl groups). Peaks at 1580 cm-1 and 1286 cm-1 indicated aromatic C=C and C-O stretching, respectively. O-H stretching (phenols) and C-H stretching (aldehydes/methylene) were observed at 3707 cm-1 and 2842 cm-1. Additional peaks at 1822 cm-1 and 1445 cm-1 were attributed to C=O stretching (cyclic ketones) and C-H stretching (alkanes). New functional groups formed during synthesis were indicated by peaks at 1822 cm-1 (C=O in cyclic ketones) and 1445 cm-1 (C-H in alkanes), with additional peaks at 1580 cm-1 and 1286 cm-1 suggesting aromatic C=C and C-O stretching. The spectra of FTIR provide clear evidence of functional group changes before and after CrS-AgNPs synthesis, as shown in Figures 4a and b. The observed peaks, corresponding functional groups, and their significance. Phenolic compounds, due to their benzene rings and alkyl chains, exhibit antimicrobial activity against multidrug-resistant bacteria by disrupting membranes through passive diffusion. Flavonoids and alkyl groups impair bacterial virulence by targeting phospholipid bilayers, while other groups like aldehydes, methylene, carboxylic acids, and nitriles interfere with membranes, nucleic acids, and metabolism. These effects are largely linked to compound lipophilicity rather than specific structures. Further research is needed to clarify the mechanisms and structure-activity relationships of these bacteriostatic agents.39,40
Figure 4. FTIR spectra of (a) Catharanthus roseus stem extract and, (b) Catharanthus roseus stem silver nanoparticles
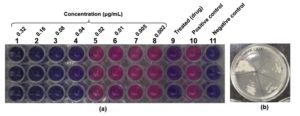
Antibacterial activity (MIC and MBC) of CrS-AgNPs against S. aureus
Bioactive phytoconstituents of plants are used as reducing and stabilizing agents in the AgNPs formation, and they possess physico-chemical characteristics such as high electric conductivity and surface area.22 The MIC of CrS-AgNPs against S. aureus was evaluated via resazurin assay. CrS-AgNPs were treated at concentrations ranging from 0.32 to 0.002 µg/mL (Figure 5), and after 18 hours of incubation, the resazurin was used to assess bacterial viability. The positive control well showed bacterial growth, while the negative and drug-treated wells did not show growth, confirming accuracy and absence of contamination. Wells from 1st to 4th demonstrated inhibition of bacterial growth (purple color), and wells from 5th to 8th indicated (turned pink) bacterial growth. Based on this observation, the MIC of CrS-AgNPs was determined to be 0.04 µg/mL. MBC against S. aureus was assessed by the streak plate method, which was performed using streak samples from the 1st to 5th wells on MHA plates and incubated for 18 hrs. The results demonstrated that wells from 2nd to 5th contained viable bacteria, while the 1st well showed complete inhibition, indicating 0.32 µg/mL MBC.
Figure 5. (a) Minimum inhibitory concentration and (b) minimum bactericidal concentration assays revealed the antibacterial efficacy of silver nanoparticles from Catharanthus roseus stem extract against the pathogenic Staphylococcus aureus
These MIC and MBC values suggest that CrS-AgNPs exhibit strong antibacterial and bactericidal activity against S. aureus at relatively low concentrations. Previous studies have reported that C. roseus methanol extracts inhibited the growth of 86.11% of multidrug-resistant Gram-negative and Gram-positive bacteria, including Escherichia coli, Pseudomonas aeruginosa, Enterobacter aerogenes, Providencia stuartii, Klebsiella pneumoniae, and Enterobacter cloacae, with MICs ranging from 64 to 1024 µg/mL.41 AgNPs synthesized using melatonin-treated C. roseus leaf inhibited the growth of S. aureus at 15 µg/ml,42 while ethanol extracts of C. roseus roots exhibited inhibition at 332 µg/mL.43 Overall, these findings demonstrate that CrS-AgNPs possess strong bactericidal potential against S. aureus, supporting their potential application as alternative antimicrobial agents.
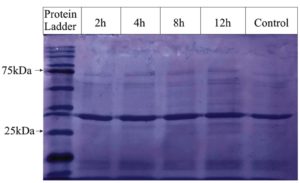
Whole-cell SDS-PAGE protein analysis
Pathogenic S. aureus was treated with CrS-AgNPs and incubated for 2, 4, 8, and 12 hours, whereas the untreated sample served as a control. Whole cell protein profiles were compared between treated and control samples using SDS-PAGE, with a protein ladder used for molecular weight reference (Figure 6). The results showed distinct changes in protein band patterns across different incubation times compared to the control, indicating variability in protein expression following CrS-AgNPs treatment in S. aureus. Some studies have reported that the ethanolic extract of C. roseus root reduces the expression of penicillin-binding protein 2a in S. aureus.44 Similarly, synthesized AgNPs using Eucalyptus camaldulensis leaf extract significantly decreased the expression of the outer membrane protein A in Acinetobacter baumannii, enhanced penetration into the bacterial cytoplasm, and disrupted intracellular processes, including DNA, RNA, and protein synthesis, as well as immune evasion.45 In addition, AgNPs-treated bacteria showed an upregulation of glutathione S-transferase and a downregulation of glutathione, superoxide dismutase, and catalase activities.46 However, due to the limitations of SDS-PAGE, further detailed analysis is needed to elucidate the precise mechanism of CrS-AgNPs action.
plc gene expression via RT-PCR
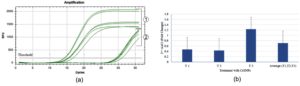
The messenger RNA levels of plc, toxicity-associated gene were examined using RT-PCR (Figure 7). plc enzyme hydrolyzes phosphatidylinositol (PI) and cleaves glycosyl-PI-linked proteins from the host cell membrane, thereby promoting bacterial survival in human blood and neutrophils.9 Gene expression analysis using RT-qPCR showed a reduction in plc transcript levels in S. aureus upon treatment with CrS-AgNPs (average fold change: 0.71 ± 0.26 (SEM)) relative to untreated controls. This suggests potential downregulation of the plc gene in response to nanoparticle exposure (~29% decrease on average). Although a downward trend in plc expression was observed in NPs-treated S. aureus, statistical analysis showed no significant variation (P = 0.438), suggesting a possible biological effect that requires further investigation. Previous studies have reported that AgNPs synthesized using aqueous extracts of Carica papaya, Cannabis sativa, and Elettaria cardamomum downregulated the expression of biofilm- and virulence-related genes such as icaC, icaD, and spa in drug-resistant S. aureus and other bacterial pathogens.47
Figure 7. Phospholipase C (plc) gene expression analysis. (a) Amplification plot showing: 1. 16S rRNA (reference gene) and 2. plc gene (target gene). (b) Bar graph representing the relative gene expression (2-ΔΔCt fold change) of plc in Catharanthus roseus stem-based silver nanoparticle-treated samples compared to control
This study introduces a modified method of synthesis of cost-effective and sustainable AgNPs using CrS aqueous extract and evaluates inhibitory concentration against pathogenic S. aureus, providing insights into molecular mechanisms and potential therapeutic applications. These plant-based AgNPs exhibited enhanced antibacterial activity against pathogenic S. aureus bacteria while altering whole-cell protein expression profiles. RT-PCR analysis further suggested a potential suppression of the virulence gene, but this requires further investigation. To the best of our knowledge, this is among the first reports to employ CrS extract specifically for the green synthesis of AgNPs and to evaluate their effect on the plc virulence gene expression in S. aureus. The findings highlight the potential of CrS-AgNPs as an alternative antibacterial agent, particularly against pathogenic S. aureus. However, further in vitro, in vivo, and clinical studies, along with stability assessments, are required to confirm their therapeutic efficacy and safety.
ACKNOWLEDGMENTS
The authors would like to express their gratitude to Maulana Azad National Institute of Technology, Bhopal, India, for providing the necessary research facilities, Dr. Bharat Kumar K. Modhera and Mr. Jamna Prasad Gujar facilitated FTIR at the Department of Chemical Engineering, and to Dr. C. Sasikumar, Materials & Metallurgical Engineering, for providing access to the XRD facilities.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
SP conceptualized and designed the study. SP performed experiments. SW contributed to the data curation. SP wrote the manuscript. SW and KMP reviewed and revised the manuscript. KMP supervised the study. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Ikuta KS, Swetschinski LR, Aguilar GR, et al. Global mortality associated with 33 bacterial pathogens in 2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet. 2022;400(10369):2221-2248.
Crossref - Mendelson M, Matsoso MP. The World Health Organization Global Action Plan for antimicrobial resistance. South African Med J. 2015;105(5):325.
Crossref - Tong SYC, Fowler Jr VG, Skalla L, Holland TL. Management of Staphylococcus aureus Bacteremia: A Review. JAMA. 2025;334(9):798-808.
Crossref - Michalik M, Podbielska-Kubera A, Dmowska-Koroblewska A. Antibiotic Resistance of Staphylococcus aureus Strains—Searching for New Antimicrobial Agents—Review. Pharmaceuticals. 2025;18(1):81.
Crossref - Cheung GYC, Bae JS, Otto M, et al. Pathogenicity and virulence of Staphylococcus aureus. Virulence. 2021;12(1):547-569.
Crossref - Prabha S, Chauhan P, Warkare S, Pandey KM. A computational investigation of potential plant-based bioactive compounds against drug-resistant Staphylococcus aureus of multiple target proteins. J Biomol Struct Dyn. 2025;43(7):3311-3329.
Crossref - Jenul C, Horswill AR. Regulation of Staphylococcus aureus Virulence. Microbiol Spectr. 2019;7(2):10.1128/microbiolspec.gpp3-0031-2018.
Crossref - Abdelmalek N, Yousief SW, Bojer MS, Alobaidallah MSA, Olsen JE, Paglietti B. The Secondary Resistome of Methicillin-Resistant Staphylococcus aureus to β-Lactam Antibiotics. Antibiot. 2025;14(2):112.
Crossref - Nakamura Y, Kanemaru K, Shoji M, et al. Phosphatidylinositol-specific phospholipase C enhances epidermal penetration by Staphylococcus aureus. Sci Rep. 2020;10:17845.
Crossref - Tiwari N, Lopez-Redondo M, Miguel-Romero L, et al. The SrrAB two-component system regulates Staphylococcus aureus pathogenicity through redox sensitive cysteines. Proc Natl Acad Sci U S A. 2020;117(20):10989-10999.
Crossref - Salam MA, Al-Amin MY, Salam MT, et al. Antimicrobial Resistance: A Growing Serious Threat for Global Public Health. Healthc. 2023;11(13):1946.
Crossref - Arunachalam K, Pandurangan P, Shi C, Lagoa R. Regulation of Staphylococcus aureus Virulence and Application of Nanotherapeutics to Eradicate S. aureus Infection. Pharmaceutics. 2023;15(2):310.
Crossref - Mendonce KC, Palani N, Rajadesingu S, Radhakrishnan K, Ayyar M, Priya LS. Pharmacological potential of bioactive compounds in Catharanthus roseus extract: A comprehensive review. Toxicol Reports. 2025;14:101998.
Crossref - Yang Y, Ding L, Zhou Y, Guo Z, Yu R, Zhu J. Establishment of recombinant Catharanthus roseus stem cells stably overexpressing ORCA4 for terpenoid indole alkaloids biosynthesis. Plant Physiol Biochem. 2023;196:783-792.
Crossref - Banyal A, Tiwari S, Sharma A, et al. Vinca alkaloids as a potential cancer therapeutics: recent update and future challenges. 3 Biotech. 2023;13(6):211.
Crossref - Jesus JA, Lago JHG, Laurenti MD, Yamamoto ES, Passero LFD. Antimicrobial activity of oleanolic and ursolic acids: an update. Evidence-based Complement Altern Med. 2015;2015:620472.
Crossref - Kumar S, Singh A, Kumar B, Singh B, Bahadur L, Lal M. Simultaneous quantitative determination of bioactive terpene indole alkaloids in ethanolic extracts of Catharanthus roseus (L.) G. Don by ultra high performance liquid chromatography–tandem mass spectrometry. J Pharm Biomed Anal. 2018;151:32-41.
Crossref - Yang L, Wang H, Zu Y gang, et al. Ultrasound-assisted extraction of the three terpenoid indole alkaloids vindoline, catharanthine and vinblastine from Catharanthus roseus using ionic liquid aqueous solutions. Chem Eng J. 2011;172(2-3):705-712.
Crossref - Tiong SH, Looi CY, Hazni H, et al. Antidiabetic and Antioxidant Properties of Alkaloids from Catharanthus roseus (L.) G. Don. Molecules. 2013;18(8):9770-9784.
Crossref - Usia T, Watabe T, Kadota S, Tezuka Y. Cytochrome P450 2D6 (CYP2D6) inhibitory constituents of Catharanthus roseus. Biol Pharm Bull. 2005;28(6):1021-1024.
Crossref - Joshi S, Huo C, Budhathoki R, et al. HPLC-ESI-HRMS/MS-Based Metabolite Profiling and Bioactivity Assessment of Catharanthus roseus. Plants. 2025;14(15):2395.
Crossref - Duman H, Eker F, Akdasci E, Witkowska AM, Bechelany M, Karav S. Silver Nanoparticles: A Comprehensive Review of Synthesis Methods and Chemical and Physical Properties. Nanomaterials. 2024;14(18):1527.
Crossref - Wahab S, Khan T, Adil M, Khan A. Mechanistic aspects of plant-based silver nanoparticles against multi-drug resistant bacteria. Heliyon. 2021;7(7):e07448.
Crossref - Nguyen NTT, Nguyen LM, Nguyen TTT, Nguyen TT, Nguyen DTC, Van Tran T. Formation, antimicrobial activity, and biomedical performance of plant-based nanoparticles: a review. Environ Chem Lett. 2022;20(4):2531-2571.
Crossref - Mostafa RG, Ali SAM, Shoueir K, Lasheen AF, Tayel AE, Sabal MS. Characterization of Virulence Genetic Profile and Potential Effect of Nanoparticles against Multidrug-Resistant Uropathogenic E. coli at Menuofia University Hospitals. J Pure Appl Microbiol. 2025;19(3):2141-2153.
Crossref - Anju S, Sarada J. Development and Evaluation of Biogenic Nano-Hydrogel Incorporating Silver Nanoparticles for Wound Healing and Antibiofilm Activity. J Pure Appl Microbiol. 2025;19(3):1938-1947.
Crossref - Restrepo CV, Villa CC. Synthesis of silver nanoparticles, influence of capping agents, and dependence on size and shape: A review. Environ Nanotechnology, Monit Manag. 2021;15:100428.
Crossref - Krishnaraj C, Jagan EG, Rajasekar S, Selvakumar P, Kalaichelvan PT, Mohan N. Synthesis of silver nanoparticles using Acalypha indica leaf extracts and its antibacterial activity against water borne pathogens. Colloids Surf B Biointerfaces. 2010;76(1):50-56.
Crossref - Pasieczna-Patkowska S, Cichy M, Flieger J. Application of Fourier Transform Infrared (FTIR) Spectroscopy in Characterization of Green Synthesized Nanoparticles. Molecules. 2025;30(3):684.
Crossref - Jesche A, Fix M, Kreyssig A, Meier WR, Canfield PC. X-Ray diffraction on large single crystals using a powder diffractometer. Philos Mag. 2016;96(20):2115-2124.
Crossref - Jahan Tamanna N, Sahadat Hossain M, Mohammed Bahadur N, Ahmed S. Green synthesis of Ag2O & facile synthesis of ZnO and characterization using FTIR, bandgap energy & XRD (Scherrer equation, Williamson-Hall, size-train plot, Monshi- Scherrer model). Results Chem. 2024;7:101313.
Crossref - Abdi RD, Dunlap JR, Gillespie BE, Ensermu DB, Almeida RA, Kerro Dego O. Comparison of Staphylococcus aureus surface protein extraction methods and immunogenicity. Heliyon. 2019;5(10):e02528.
Crossref - Prabha S, Warkare S, Ranga A, Pandey KM. Exploring the Protein Targets of Cinnamomum zeylanicum’s Phytoconstituents Against Pathogenic Staphylococcus aureus: GC-MS Profiling, Molecular Docking, Pharmacophore Modeling, and Pathway Analysis. Trop J Nat Prod Res. 2025;9(10):5061-5072.
Crossref - Bhusal M, Pathak I, Bhadel A, Shrestha DK, Sharm KR. Synthesis of silver nanoparticles assisted by aqueous root and leaf extracts of Rhus chinensis Mill and its antibacterial activity. Heliyon. 2024;10(13):e33603.
Crossref - Vivek-Ananth RP, Mohanraj K, Sahoo AK, Samal A. IMPPAT 2.0: An Enhanced and Expanded Phytochemical Atlas of Indian Medicinal Plants. ACS Omega. 2023;8(9):8827-8845.
Crossref - Indana MK, Gangapuram BR, Dadigala R, Bandi R, Guttena V. A novel green synthesis and characterization of silver nanoparticles using gum tragacanth and evaluation of their potential catalytic reduction activities with methylene blue and Congo red dyes. J Anal Sci Technol. 2016;7(1):1-9.
Crossref - Balashanmugam P, Kalaichelvan PT. Biosynthesis characterization of silver nanoparticles using Cassia roxburghii DC. aqueous extract, and coated on cotton cloth for effective antibacterial activity. Int J Nanomedicine. 2015;10(Suppl 1):87-97.
Crossref - Adil M, Alam S, Amin U, et al. Efficient green silver nanoparticles-antibiotic combinations against antibiotic-resistant bacteria. AMB Express. 2023;13(1):115.
Crossref - Lobiuc A, Paval NE, Mangalagiu II, et al. Future Antimicrobials: Natural and Functionalized Phenolics. Molecules. 2023;28(3):1114.
Crossref - Ecevit K, Barros AA, Silva JM, Reis RL. Preventing Microbial Infections with Natural Phenolic Compounds. Futur Pharmacol. 2022;2(4):460-498.
Crossref - Voukeng IK, Beng VP, Kuete V. Antibacterial activity of six medicinal Cameroonian plants against Gram-positive and Gram-negative multidrug resistant phenotypes. BMC Complement Altern Med. 2016;16(1):388.
Crossref - Sheshadri SA, Sriram S, Balamurugan P, et al. Melatonin improves bioreductant capacity and silver nanoparticles synthesis using Catharanthus roseus leaves. RSC Adv. 2015;5(59):47548-47554.
Crossref - Shil A, Banerjee A, Maji BK, Bishayi B, Sikdar (ne’e Bhakta) M. Multiple antibiotic resistant Staphylococcus aureus induced hepatocellular anomaly: A possible amelioration by Catharanthus roseus (L.) G.Don. South African J Bot. 2022;148:446-459.
Crossref - Shil A, Mukherjee S, Biswas P, et al. Catharanthus roseus (L.) G. Don counteracts the ampicillin resistance in multiple antibiotic-resistant Staphylococcus aureus by downregulation of PBP2a synthesis. Open Life Sci. 2023;18(1):20220718.
Crossref - Syukri DM, Singh S, Mohite P, et al. A Novel Anti-Virulence Approach For Attenuating Acinetobacter Baumannii Outer Membrane Protein A Using Biogenic Silver Nanoparticles: In Vitro and In Silico Docking. Regen Eng Transl Med. 2025.
Crossref - Yuan YG, Peng QL, Gurunathan S. Effects of silver nanoparticles on multiple drug-resistant strains of Staphylococcus aureus and Pseudomonas aeruginosa from mastitis-infected goats: An alternative approach for antimicrobial therapy. Int J Mol Sci. 2017;18(3):569.
Crossref - Singh J, Agrawal RK, Bankoti K, et al. Antibacterial, anti-biofilm and anti-virulence activity of biosynthesized silver nanoparticles against drug-resistant Staphylococcus aureus. Vet Res Commun. 2025;49(6):345.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.