ISSN: 0973-7510
E-ISSN: 2581-690X
Paracetamol is a widely recognized pharmaceutical contaminant in the environment, emphasizing the need for effective biotransformation strategies for its remediation. This study explored the biotransformation ability of Alcaligenes sp., isolated from waste-burned soil, using high performance liquid chromatography analysis. Bacterial isolation was performed on nutrient agar containing 50 mg/L paracetamol as the sole carbon source, followed by identification through Gram staining and ALSP testing. Paracetamol biotransformation was conducted in liquid nutrient medium and incubated at 30 °C on a shaker for 72 hours. Three bacterial isolates (RZP1, RZP2, and RZP3) were obtained, with RZP2 and RZP3 confirmed as Alcaligenes sp. All isolates were Gram-negative and exhibited distinct morphological characteristics. Thin layer chromatography showed varying Rf values for each isolate: RZP1 (0.48), RZP2 (0.58), and RZP3 (0.74), compared to 0.70 for the paracetamol standard. HPLC analysis revealed a complex biotransformation profile, with five metabolite peaks detected at 1.657, 2.418, 2.975, 5.311, and 8.225 minutes, whereas the paracetamol standard exhibited a single peak at 5.71 minutes. The total metabolite peak area reached 76,184,224, indicating extensive biotransformation involving both hydrolytic and oxidative pathways. These results demonstrate that Alcaligenes sp. can biotransform paracetamol into multiple metabolites with diverse polarity profiles, highlighting its potential application in environmentally sustainable pharmaceutical bioremediation.
Paracetamol, Biotransformation, Alcaligenes sp., HPLC, Drug Metabolites
Biotechnology has experienced rapid development,1 and the increasing public demand for sustainable pharmaceutical products has positioned biotechnology as a key solution for meeting these needs while addressing environmental challenges.2 However, the rapid growth of the pharmaceutical industry has negatively impacted the environment, particularly through the discharge of pharmaceutical compounds into aquatic and terrestrial ecosystems.3
Acetaminophen (APAP), widely known as paracetamol, is among the most commonly used analgesic and antipyretic drugs globally for managing headaches, fever, and mild pain.4 Its widespread availability and low cost contribute to an estimated annual production of approximately 145,000 tons worldwide.5 This extensive consumption makes paracetamol a significant environmental concern, as it is frequently detected in various environmental matrices, including wastewater, surface water, and groundwater.6-8 The increasing prevalence of pharmaceutical pollutants underscores the need for effective biotechnological approaches for their transformation and remediation.9
Microbial biotransformation has emerged as a promising strategy for processing pharmaceutical compounds due to the enzymatic specificity that enables selective and efficient substrate conversion.10 Biological waste treatment relies on microorganisms such as bacteria, fungi, and protozoa to break down organic waste into simpler compounds, converting them into CO2‚ H2O, and biomass that support microbial growth and reproduction.11 Microorganisms also play critical roles in food and pharmaceutical production.12 Previous studies have reported successful biotransformation of bioactive compounds using endophytic and soil microbial isolates, such as the conversion of catechins from gambir plants (Uncaria gambir Roxb.) into novel metabolites with diverse bioactivities.13
Soil represents a natural habitat rich in microbial diversity, particularly bacteria.14,15 As a component of the earth’s crust composed of minerals and organic matter, soil provides nutrients and water essential for microbial survival.16 Waste-burned soil contains abundant minerals and organic compounds, making it both a fertile growth medium and a promising source of metabolically versatile bacteria.17 The high cellulose content in landfill soil, derived from organic materials such as leaves and household waste, creates a natural environment for cellulolytic bacteria.18 Bacterial groups capable of cellulose hydrolysis include Achromobacter, Angiococcus, Bacillus, Cellulomonas, Cytophaga, Clostridium, Cellivibrio, Flavobacterium, Pseudomonas, Poliangium, Sorangium, Sporocytophaga, Vibrio, and Cellfalcicula,19 as well as Citrobacter, Serratia, Klebsiella, Enterobacter, and Aeromonas.20
Among these microbial groups, the genus Alcaligenes has been identified as capable of degrading various aromatic compounds.21 Studies indicate that Alcaligenes sp. isolated from activated sludge can degrade indole through metabolic pathways involving indoxyl, isatin, anthranilic acid, and gentisic acid.22 This metabolic versatility suggests that Alcaligenes species possess enzymatic systems suitable for the biotransformation of pharmaceutical compounds such as paracetamol. High performance liquid chromatography (HPLC) has become the analytical standard for characterizing biotransformation products because of its high resolution, sensitivity, and reproducibility.23 Its application enables precise monitoring of substrate degradation and metabolite formation, providing essential insights into microbial metabolic pathways. Despite growing interest in microbial biotransformation of pharmaceutical compounds, research exploring bacteria from unique environments such as waste burned soil remains limited. Such environments may harbor novel bacterial strains with unique metabolic capabilities relevant to pharmaceutical biotransformation.
Based on this background, this study aimed to investigate the biotransformation of paracetamol by Alcaligenes sp. isolated from waste burned soil and to identify the resulting metabolite profile using HPLC analysis. The findings are expected to contribute valuable knowledge to advance biotechnology applications in pharmaceutical processing and environmental remediation.
Chemicals and growth medium
Paracetamol (C8H9 NO2, 151.17 g/mol) was obtained in analytical grade purity from Sigma-Aldrich (St. Louis, MO, USA). Nutrient agar (NA; Merck, Darmstadt, Germany), nutrient broth (NB; HiMedia, Mumbai, India), and all analytical grade reagents, including PA grade methanol (Fisher Scientific, USA), were used throughout the study.
Bacterial growth and paracetamol transformation
The study employed a laboratory based qualitative descriptive method.24 Soil samples from burned waste were collected aseptically from a waste incineration site in the Sarojo area, Bukittinggi City, West Sumatra. Two grams of soil (Sample ID: S1) were transferred into sterile containers, stored in a cool box, and transported to the laboratory. Bacteria were isolated following standard soil dilution plating procedures on nutrient agar.25 One gram of soil was added to a sterile test tube containing 10 mL nutrient broth and homogenized using a vortex mixer for 30 seconds. Serial dilutions were performed up to 10-1. From the 10-1 to 10-1 dilutions, 1 mL of each suspension was plated onto sterile Petri dishes, followed by the addition of 15 mL molten NA supplemented with 50 mg/L paracetamol as the sole carbon source. Plates were gently swirled and incubated at 37 °C for 48 hrs. Distinct colonies were purified using the quadrant streak method and incubated again at 37 °C for 48 hrs. Colony morphology, including pigmentation, form, elevation, and margin, was systematically recorded following standard microbiological guidelines.26
Biotransformation assays were conducted using 15 mL sterile nutrient broth in culture bottles sterilized via autoclaving at 121 °C for 15 min at 2 atm. After cooling, selected isolates were inoculated using two sterile inoculating needles and incubated for 24 hrs at 30 °C and 150 rpm to obtain pre cultures. A 50 mg/L paracetamol solution was then added, and cultures were incubated at 30 °C and 150 rpm for 72 hrs. A control sample consisting of medium and paracetamol without bacterial inoculation was prepared in parallel. After incubation, cultures were centrifuged at 6,000 rpm for 20 min. The supernatant was filtered through a 0.22 µm syringe filter (Millipore, USA) prior to solvent extraction with PA grade methanol. Extracts were analyzed using thin layer chromatography (TLC), preparative TLC (TLCP), and HPLC.
TLC and TLCP analysis
TLC was performed on silica gel 60 F254 plates (Merck, Germany) using a methanol:chloroform (7:3 v/v) mobile phase according to established analytical protocols. Plates were developed in a saturated chamber for 20 minutes and visualized under UV light at 254 nm. For TLCP, silica bands were scraped, eluted with methanol, vortexed, and filtered prior to HPLC analysis.27
HPLC analysis
HPLC was conducted using a Shimadzu LC-20AT system equipped with UV Vis detection and LabSolutions software. Separation was performed on a C18 column (250 mm × 4.6 mm, 5 µm; Agilent Technologies). The mobile phase consisted of methanol:water (60:40 v/v) delivered at 1.0 mL/min. The injection volume was 20 µL, the column temperature was maintained at 30 °C, and detection was carried out at 243 nm. Analytical procedures and performance characteristics followed ICH Q2 (R2) validation guidelines for specificity, precision, and reproducibility.28
Macroscopic and microscopic characterization
The macroscopic evaluation of bacterial colonies isolated from waste burned soil revealed considerable morphological variation, reflecting ecological adaptation to environments enriched with thermally modified organic residues and xenobiotic by products. Post combustion soil systems typically exhibit altered pH, residual combustion hydrocarbons, and structurally oxidized substrates, which act as selective agents favoring thermotolerant, stress resistant, and xenobiotic metabolizing microorganisms. The three isolates displayed distinct visual and microscopic characteristics. The macroscopic characteristics of the bacterial isolates are summarized in Table 1. Colonies were evaluated based on six categories: color, shape, edge, elevation, size, and texture.
Table (1):
Macroscopic characteristics of bacterial isolates from soil burned waste
Isolate |
Color |
Shape |
Edge |
Elevation |
Size |
Texture |
|---|---|---|---|---|---|---|
RZP1 |
White |
Irregular |
Undulate |
Flat |
Large |
Rough |
RZP2 |
White |
Circular |
Entire |
Convex |
Moderate |
Smooth |
RZP3 |
White |
Circular |
Entire |
Convex |
Small |
Smooth |
Macroscopic characteristics were recorded based on six categories: color, shape, edge, elevation, size, and texture of bacterial colonies.29 Colony morphology was documented in terms of pigmentation, elevation, optical properties, margin integrity, and surface texture, following established microbiological classification frameworks. These variations are essential indicators of adaptive enzyme system expression and resource acquisition strategies, often correlating with metabolic specialization. Observed colony phenotypes showed clear distinctions: RZP1 exhibited an irregular and wavy morphology with a rough texture and large size, whereas RZP2 and RZP3 formed round colonies with intact edges and convex elevations but differed in size and texture.30 Morphological differentiation among isolates strongly suggests divergent metabolic competencies relevant to biotransformation.
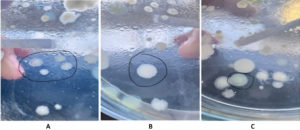
Sukmawati and Rosalina31 reported that two bacterial isolate species from soil can inhibit the growth of pathogenic bacteria by producing antibiotic compounds.32 Microorganisms’ ability to detoxify xenobiotic compounds allows them to survive in toxic environments, utilizing carbon, phosphorus, sulfur, and nitrogen from available sources. Diverse microbial populations in harsh, contaminated environments demonstrate significant potential to degrade and transform pollutants. Waste-incineration environments contain a variety of complex organic compounds, including pharmaceutical residues, plastics, and other chemicals, creating high selective pressure on microorganisms. The persistence of emerging contaminant compounds (CECs) in the environment has led to the evolution of microbes capable of utilizing them as a sole source of carbon and energy. These conditions favor the natural selection of bacteria with specialized enzymatic systems for the degradation of xenobiotic compounds such as paracetamol. Distinct macroscopic colony morphologies were observed among the isolates (Figure 1A-C). Isolate RZP1 (Figure 1A) exhibited irregular colonies with undulate margins and a rough surface texture. In contrast, RZP2 (Figure 1B) formed circular colonies with smooth surfaces, entire margins, and convex elevation. Similarly, RZP3 (Figure 1C) displayed circular colonies with convex elevation, although slight variations in size and surface characteristics were observed compared to RZP2.
Figure 1. Macroscopic colony morphology of bacterial isolates from waste-burned soil. (A) Isolate RZP1 exhibiting irregular colony shape with undulate margins and rough surface texture. (B) Isolate RZP2 displaying circular colonies with smooth surface, entire margins, and convex elevation. (C) Isolate RZP3 showing circular colonies with convex elevation and distinct size variation compared to RZP2
Gram staining and identification
Gram staining revealed a uniform Gram-negative reaction among all isolates, consistent with Alcaligenes related taxa, which are characterized by thin peptidoglycan layers and outer membranes rich in lipopolysaccharides. The Gram-negative profile is strongly associated with environmental resilience, especially in soils contaminated with combustion derived aromatic compounds. Such bacteria typically possess outer membrane efflux systems, permeability barriers, and inducible catabolic enzymes, enabling survival and substrate utilization in chemically stressed habitats. The detailed Gram staining results are summarized in Table 2. Subsequent biochemical identification using the ALSP test further confirmed that isolates P2 and P3 were affiliated with Alcaligenes sp., while P1 showed a negative reaction, as presented in Table 3.
Table (2):
Gram staining results of bacterial isolates from Waste-Burned Soil
No. |
Sample Origin |
Sample Owner |
Batch Code |
Test Results |
|---|---|---|---|---|
1 |
Location 1 |
Mhd Riza Marjoni |
P1.1 |
Damaged |
2 |
Location 2 |
Mhd Riza Marjoni |
P2.2 |
Gram-negative |
3 |
Location 3 |
Mhd Riza Marjoni |
P3.2 |
Gram-negative |
Table (3):
Bacterial Isolation and Identification
No. |
Sample Origin |
Sample Owner |
Batch Code |
ALSP Test Results |
|---|---|---|---|---|
1 |
Location 1 |
Mhd Riza Marjoni |
P1 |
– |
2 |
Location 2 |
Mhd Riza Marjoni |
P2 |
+ |
3 |
Location 3 |
Mhd Riza Marjoni |
P3 |
+ |
Description: ALSP: Alcaligenes sp.; (+): Positive; (-): Negative
TLC profile of paracetamol and metabolites
TLC successfully distinguished paracetamol from its biotransformation products. The paracetamol standard consistently exhibited an Rf value of 0.70, whereas metabolite bands exhibited Rf values ranging from 0.48-0.74, reflecting shifts in compound polarity due to microbial enzymatic modifications such as hydroxylation, deacetylation, or partial ring oxidation. These variations indicate that the isolates possess diverse enzymatic functionalities capable of modifying multiple structural regions of the parent compound. Such chemical behavior aligns with documented Alcaligenes associated xenobiotic degradation pathways, where microbes express multifunctional oxygenases and hydrolases to process complex substrates.
The distribution of chromatographic mobility values for paracetamol and its metabolites is summarize in the Table 4.
Table (4):
Results of Rf Value Measurements
Standard and Sample |
Spot Height |
Rf value |
|---|---|---|
Paracetamol |
5.6 |
0.7 |
RZP-1 |
3.2 |
0.48 |
RZP-2 |
4.6 |
0.58 |
RZP-3 |
3.8 |
0.74 |
HPLC analysis
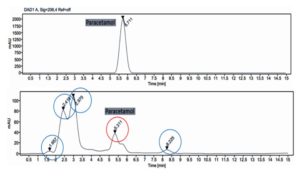
HPLC provided definitive confirmation of paracetamol biotransformation. The paracetamol standard consistently eluted at 5.71 minutes under validated chromatographic conditions, fully meeting ICH Q2 (R2) specificity requirements. In contrast, the biotransformation extract exhibited four distinct metabolite peaks at 2.418, 2.975, 5.311, and 8.225 minutes, each representing a unique intermediate or end product in the degradation pathway.
The corrected total peak area of 7618.8254 quantitatively reflects metabolic turnover. Early-eluting peaks indicate increased polarity due to hydroxylation or deacetylation, whereas later-eluting peaks suggest retained hydrophobicity or partial aromatic ring integrity. These retention behaviors support the occurrence of multi-stage enzymatic transformations. The HPLC chromatogram is presented in Figure 2.
The detailed DAD-A peak parameters recorded at 206.4 nm, including retention time, area, and peak characteristics, are summarized in Table 5.
Table (5):
DADI A Spectroscopy Data (Sig = 206.4 Ref = off)
Peak |
Time (RT) [min] |
Area |
Height |
Tf |
|---|---|---|---|---|
1 |
2.418 |
2284.9822 |
1248.11 |
0.77069 |
2 |
3.975 |
3,628.7741 |
1.93 |
2.419911 |
3 |
5.311 |
1433.2421 |
– |
0.95389 |
4 |
8.225 |
186.1618 |
2.44 |
0.92222 |
Total |
7618.8254 |
Proposed biotransformation pathways
Integration of TLC and HPLC data suggests that paracetamol underwent a sequence of microbial transformations, likely involving hydroxylation, deacetylation, and oxidative cleavage. Hydroxylation reduces Rf values by increasing polarity, clarifying previous misinterpretations. These pathways align with biochemical models reported for Alcaligenes species, which express broad specificity monooxygenases, amide hydrolases, and ring cleavage dioxygenases. This metabolic versatility highlights their potential utility in environmental bioremediation targeting pharmaceutical pollutants.33
Sustainability Assessment (Green Analytical Chemistry [GAC] and White Analytical Chemistry [WAC])
The GAC scores were assigned based on the 12 established principles of Green Analytical Chemistry, each evaluated according to the characteristics of the analytical workflow used in this study.34 The scoring scale ranged from 0 to 4, where 0 indicates poor alignment with sustainability principles and 4 indicates excellent adherence. Several factors influenced the scoring.
Solvent usage and toxicity
Only methanol (PA grade) was used as the organic solvent and in relatively low volumes. Because methanol presents moderate toxicity, parameters related to solvent safety and toxicity were assigned mid-range scores (2-3).
Waste generation
The method produced low amounts of waste due to small HPLC solvent flow rates and minimal solid waste from TLC plates. This justified higher scores (3).
Energy consumption
Microbial incubation at 30 °C and HPLC analysis at room temperature required minimal energy input, supporting greener scoring (3).
Safety considerations
The absence of corrosive reagents, strong acids, strong bases, or carcinogenic chemicals resulted in a score of 4 for operator safety.
Sample preparation design
As no derivatization or multi-step purification was required, the method scored highly for minimal sample preparation.
Material reuse and life-cycle considerations
Although the HPLC column is reusable, solvents cannot be recycled, which reduced the score in this category (2).
Overall, the total GAC score reflects a balanced methodology that is green, safe, and efficient, although still limited by reliance on methanol and non-recyclable solvents. This systematic scoring ensures transparency and supports reproducibility of the assessment. The analytical procedure aligns with the principles of Green Analytical Chemistry by prioritizing reduced solvent usage, minimal reagent toxicity, and efficient chromatographic operation. The full GAC scoring framework, evaluating greenness, operational safety, and material efficiency, should be provided in Table 6.
Table (6):
Green Analytical Chemistry (GAC) Evaluation
No. |
GAC Principle |
Score (0-4) |
Justification |
|---|---|---|---|
1 |
Minimization of solvent use |
3 |
Low-volume methanol used for extraction and HPLC; no large-scale solvent consumption. |
2 |
Use of safer solvents and reagents |
3 |
Methanol PA is the only organic solvent; no chlorinated or highly toxic solvents involved. |
3 |
Waste reduction |
3 |
Liquid waste from HPLC is limited, and solid waste from TLC plates is minimal. |
4 |
Reduction of analytical toxicity |
2 |
Methanol has moderate toxicity but no corrosive or carcinogenic reagents were used. |
5 |
Energy efficiency |
3 |
Incubation at 30 °C and HPLC at room temperature require low energy input. |
6 |
Instrument miniaturization/low-impact instruments |
3 |
Standard HPLC with low solvent flow rates; no high-energy equipment. |
7 |
Operator safety |
4 |
No highly hazardous chemicals; standard microbiology and chromatography operations. |
8 |
Minimal sample preparation |
3 |
No derivatization; workflow uses direct extraction and straightforward chromatography. |
9 |
Overall environmental friendliness |
3 |
Method avoids environmentally hazardous reagents and minimizes waste. |
10 |
Material reuse/recycling |
2 |
HPLC columns reusable; solvent reuse limited. |
11 |
Reduction of consumables |
3 |
Limited use of TLC plates, vials, and culture tubes. |
12 |
Overall greenness of the method |
3 |
Technique aligns well with sustainable analytical practices. |
Total GAC Score = 34/48 (Green Category)
The WAC evaluation was performed using the Red-Green-Blue (RGB) model, which balances analytical performance, environmental friendliness, and operational practicality.35 A score of 0-4 was assigned for each parameter based on predefined criteria, and the results are summarized in Table 7.
Table (7):
White Analytical Chemistry (WAC) Evaluation
| A. Analytical Performance (Red Criteria) | ||
|---|---|---|
| Parameter | Score (0-4) | Justification |
| Accuracy | 4 | HPLC produced sharp, well-defined peaks with validated RT (5.71 min). |
| Precision | 3 | Reproducible peak retention times and consistent metabolite profile. |
| Selectivity/Specificity | 4 | TLC + HPLC provided clear discrimination between paracetamol and metabolites. |
| Sensitivity | 3 | Adequate detection at mg/L concentration; sufficient for biotransformation studies. |
| Total Red Score | 14/16 | Excellent analytical performance. |
| B. Environmental Impact (Green Criteria) | ||
| Parameter | Score | Justification |
| Solvent toxicity | 2 | Methanol is moderately toxic but manageable. |
| Energy consumption | 3 | Low-temperature incubation and ambient-temperature chromatography. |
| Waste generation | 3 | Low volumes of organic waste and minimal solid waste. |
| Operator safety | 4 | No corrosive, carcinogenic, or highly hazardous chemicals. |
| Total Green Score | 12/16 | Low environmental impact. |
| C. Practicality (Blue Criteria) | ||
| Parameter | Score | Justification |
| Cost efficiency | 2 | HPLC operational cost moderately high. |
| Time efficiency | 3 | Workflow efficient; 72 hour culture + rapid chromatographic screening. |
| Operational simplicity | 3 | Standard microbiology + chromatography; no specialized instrumentation. |
| Reproducibility | 4 | Procedure easily replicable across laboratories. |
| Total Blue Score | 12/16 | High practicality and feasibility. |
Overall WAC Score = 38/48 (Balanced White Method)
Analytical Performance (Red Criteria)
Accuracy (score 4)
The HPLC method produced a sharp and well-resolved paracetamol peak at a validated retention time of 5.71 min, fulfilling ICH Q2 (R2) specificity requirements.
Precision (score 3)
Replicate chromatograms showed consistent retention times and reproducible metabolite profiles. Minor natural variability in microbial metabolism prevented a maximum score.
Selectivity/Specificity (score 4)
Combined TLC and HPLC analyses confirmed clear separation of parent paracetamol from its metabolites, justifying the highest score.
Sensitivity (score 3)
The analytical system successfully detected metabolites derived from 50 mg/L paracetamol, demonstrating adequate sensitivity for biotransformation studies.
These scores demonstrate strong analytical performance.
Environmental Impact (Green Criteria)
Solvent toxicity (score 2)
The use of methanol, although moderate in toxicity, limited the score.
Energy consumption (score 3)
Both microbial incubation and chromatographic analysis operated under low-energy conditions.
Waste generation (score 3)
Limited organic solvent waste and minimal solid waste justified a favorable score.
Operator safety (score 4)
The absence of corrosive, explosive, or high-risk chemicals resulted in excellent safety conditions.
These parameters align with the environmental dimension of WAC.
Practicality (Blue Criteria)
Cost efficiency (score 2)
Routine HPLC operating costs, including solvent consumption and column usage, remained moderate.
Time efficiency (score 3)
Although microbial incubation required 72 hours, analytical steps were rapid, supporting a mid-to-high score.
Operational simplicity (score 3)
The workflow relied on standard microbiological and HPLC equipment widely available in analytical laboratories.
Reproducibility (score 4)
Use of common materials and standardized protocols facilitated straightforward replication.
The WAC scoring therefore reflects a balanced and realistic evaluation of methodological practicality.
General principles used in scoring determination
Across both GAC and WAC frameworks, the following academic criteria guided score assignment.
Hazard vs. risk distinction
Assessments considered both toxicological classification and actual operator exposure risk.
Volume-based environmental assessment
Lower solvent volumes were directly associated with reduced environmental burden.
Energy-demand analysis
All procedures were evaluated based on heating, cooling, pressurization, and instrument run times.
Analytical necessity
Steps such as the combined TLC–HPLC approach were evaluated based on analytical justification rather than procedural complexity.
Interlaboratory reproducibility
Techniques employing common consumables and equipment received higher scores.
Holistic balance (WAC RGB model)
Trade-offs between sustainability and analytical performance were explicitly acknowledged.
Similarly, WAC scoring emphasizes methodological balance across analytical performance, environmental footprint, and resource expenditure. The complete WAC scoring breakdown is provided below.
This indicates strong analytical performance balanced with environmental and operational feasibility.
Summary of findings
Overall, the results demonstrate that Alcaligenes sp. isolated from waste-burned soil exhibits substantial paracetamol biotransformation capacity, generating multiple metabolites with distinct chromatographic signatures. The observed chromatographic shifts suggest the involvement of multi-step enzymatic pathways, including deacetylation, hydroxylation, and partial aromatic ring transformations, consistent with previously reported microbial degradation mechanisms.36 In addition, the combined TLC-HPLC workflow proved reliable and reproducible for monitoring biotransformation dynamics, supporting its applicability for routine analytical assessment.37 These findings align with recent studies highlighting the strategic role of metabolically versatile bacteria in sustainable, microbially driven pharmaceutical bioremediation systems.38
This study provides comprehensive evidence of the paracetamol biotransformation capacity of bacterial isolates obtained from waste burned soil, with two dominant isolates (RZP2 and RZP3) conclusively identified as Alcaligenes sp. Through integrated TLC and HPLC analyses, a diverse set of biotransformation products was characterized, including five distinct metabolites with unique chromatographic and polarity profiles, confirming the occurrence of multi-step hydrolytic and oxidative transformations. These results reinforce the role of xenobiotic adapted soil bacteria as potent agents for pharmaceutical degradation.
The chromatographic differences observed between the parent compound and its metabolites, particularly the broadened retention time distribution and altered Rf values, provide strong analytical evidence of enzymatic modifications. The findings support the involvement of deacetylation, hydroxylation, and partial aromatic ring transformation pathways within the isolates. Such mechanistic insights expand understanding of microbial paracetamol degradation and highlight the ecological relevance of microbes inhabiting post combustion environments.
Importantly, this study emphasizes the biotechnological potential of waste burned soil as a reservoir of metabolically versatile microorganisms capable of transforming pharmaceutical contaminants. The ability of Alcaligenes sp. to convert paracetamol into multiple downstream metabolites positions this genus as a promising candidate for environmentally sustainable bioremediation applications. This supports the development of biological treatment strategies that reduce reliance on chemical inputs and energy intensive processes.
From an analytical perspective, the combined TLC–HPLC workflow demonstrated high reliability, sensitivity, and reproducibility in tracking biotransformation dynamics. This approach offers a cost effective and scalable analytical framework suitable for routine monitoring, and future integration with advanced analytical techniques is expected to enable detailed structural elucidation of metabolites and enhance method robustness for larger scale applications. Overall, the findings contribute substantially to the fields of microbial biotechnology and environmental remediation by demonstrating that contaminated soils can serve as valuable sources of microorganisms with significant xenobiotic modifying capabilities.
ACKNOWLEDGMENTS
The authors would like to thank the Ministry of Education, Culture, Research, and Technology for providing research funding.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This study was funded by the Ministry of Education, Culture, Research, and Technology for providing research funding (Contract No.: 131/C3/DT.05.00/PL/2025).
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Wasilah U, Rohimah S, Su’udi, M. The Development of Biotechnology In Indonesia. Rekayasa. 2019;12:85-90.
Crossref - Pessoa MG, Vespermann KAC, Paulino BN, Barcelos MCS, Pastore GM, Molina G. Newly isolated microorganisms with potential application in biotechnology. Biotechnol Adv. 2019;37(2):319-339.
Crossref - Baroka IU. Pharmaceutical waste management: sources, environmental impacts, and sustainable solutions. Pharmacy Reports. 2025;5:95.
Crossref - Tripathi YB, Tiwari O, Nagwani S, Mishra B. Pharmacokinetic-interaction of Vitex negundo Linn. & paracetamol. Indian J Med Res. 2009;130(4):479-483.
- Sebastine IM, Wakeman RJ. Consumption and Environmental Hazards of Pharmaceutical Substances in the UK. Process Saf Environ Prot. 2003;81:229-235.
Crossref - Buxton HT, Kolpin DW. Pharmaceuticals, Hormones, and Other Organic Wastewater Contaminants in US Streams. Water Encyclopedia. 2004:605-608.
Crossref - Li J, Ye Q, Gan J. Degradation and transformation products of acetaminophen in soil. Water Res. 2014;49:44-52.
Crossref - Sousa JCG, Ribeiro AR, Barbosa MO, Pereira MFR, Silva AMT. A review on environmental monitoring of water organic pollutants identified by EU guidelines. J Hazard Mater. 2018;344:146-162.
Crossref - Wang X, Pehkonen SO, Ray AK. Photocatalytic reduction of Hg(II) on two commercial TiO2 catalysts. Electrochim Acta. 2004;49(9-10):1435-1444.
Crossref - Syahrani A, Panjaitan TS, Indrayanto G, Wilkins AL. N-Acetylation and N -Formylation of m -Aminobenzoic acid by Cell Suspension Cultures of Solanum Laciniatum. J Asian Nat Prod Res. 2000;2(4):305-309.
Crossref - Firdus, Muchlisin ZA. Degradation rate of sludge and water quality of septic tank (water closed) by using starbio and freshwater catfish as biodegradator. Jurnal Natural. 2010;10:1-6.
- Kästner, M., Miltner, A. Application of compost for effective bioremediation of organic contaminants and pollutants in soil. Appl Microbiol Biotechnol. 2016;100:3433–3449.
Crossref - Nandika D, Syamsu K, Arinana A, Kusumawardhani DT, Fitriana Y. Bioactivities of catechin from gambir (Uncaria gambir Roxb.) against wood-decaying fungi. BioRes. 2019;14(3):5646-5656.
Crossref - Oorts K, Smolders E, Lanno R, Chowdhury MJ. Bioavailability and Ecotoxicity of Lead in Soil: Implications for Setting Ecological Soil Quality Standards. Environ Toxicol Chem. 2021;40(7):1948-1961.
Crossref - Purba JM, Prasetya B. Exploration and Identification of Mycorrhizae from Various Types of Vegetation in Agroforestry. Jurnal Tanah dan Sumberdaya Lahan. 2021;8(2):369-376.
Crossref - Fajri JA, Nurmiyanto A, Sa’adah NN, et al. Selection of endophyte and indigenous bacteria degrading textile wastewater in floating treatment wetland. Int J Environ Sci Technol. 2025;22:319-330.
Crossref - Satwika TD, Yulianti DM, Hikam AR. Characteristics and enzymatic potential of bacteria from kitchen waste soil and livestock manure as candidates for organic waste biodegradation agents. Al-Hayat: Journal of Biology and Applied Biology. 2021;4:11-18.
Crossref - Ibrahim SNMM, Amalia SE, Agatha B, et al. Screening and identifying of cellulolytic bacteria from Alas Purwo National Park. Inventing prosperous future through biological research and tropical biodiversity management: Proceedings of the 5th International Conference on Biological Science. AIP Conference Proceedings. 2018;2002:020064.
Crossref - Syamsia, Kuswinanti T, Syam’un E, Masniawati A. The Potency of Endophytic Fungal Isolates Collected from Local Aromatic Rice as Indole Acetic Acid (IAA) Producer. Procedia Food Sci. 2015;3:96-103.
Crossref - Forsythe SJ, Abbott SL, Pitout J. Klebsiella, Enterobacter, Citrobacter, Cronobacter, Serratia, Plesiomonas, and Other Enterobacteriaceae. Manual of Clinical Microbiology. 2015:714-737.
Crossref - Pandey R, Swamy KV, Khetmalas MB. Indole: A novel signaling molecule and its applications. Indian J Biotechnol. 2013;12(3):297-310.
- Chen X, Shen W, Chen J, Zhu Y, Chen C, Xie S. Tetracycline biotransformation by a novel bacterial strain Alcaligenes sp. T17. Sci Total Environ. 2022;832:155130.
Crossref - Lestari PED, Jannah AKR, Hisan HK, Fitri MN, Azharani NA. Comparative analysis of HPLC and other techniques for antibiotics detection. Termometer: Jurnal Ilmiah Ilmu Kesehatan dan Kedokteran. 2023;2(2):01-11.
Crossref - de Oliveira EMS, Freitas JB. Assessment of the Role of Laboratory Facilities in Science Learning: A Qualitative Study of Pre-secondary School in Uailili, Baucau, Timor-Leste. Asian Journal of Education and Social Studies. 2025;51(12):70-85.
Crossref - Kaufman DD, Williams LE, Sumner CB. Effect of plating medium and incubation temperature on growth of fungi in soil-dilution plates. Can J Microbiol. 1963;9(6):741-751.
Crossref - Hatagale VD, Somwanshi SD, Pawar SV. Effect of Culture Media on Cultural and Morphological Characteristics of Sclerotium rolfsii. J Sci Res Rep. 2025;31(10):578-584.
Crossref - Zych M, Pyka-Pajak A. TLC in the Analysis of Plant Material. Processes. 2025;13(11):3497.
Crossref - Peikova L, Koleva P, Balkanski S, Valtchev V, Choparinova P. Methodology for the analysis of etomidate and its enantiomer by HPLC. Pharmacia. 2025;72:1-7.
Crossref - Khalila R, Fitri L, Suhartono S. Isolation and Characterization of Thermophilic Bacteria as Cellulolytic Enzyme Producer from the Hot Spring of Ie Seuum Aceh Besar, Indonesia. Microbiol Indones. 2020;14(1):4.
Crossref - Syarifahanum S, Manalu K, Nasution RA. Isolation and test of the ability of fungi to degredate polyethylene plastic from landfill plunges. Jurnal Pembelajaran Biologi: Kajian Biologi dan Pembelajarannya. 2024;11(1):22-34.
Crossref - Sukmawati S, Rosalina F. Antimikrob Bacteria Isolation from Soils as Producing Antimicrob Compounds. Biospecies. 2020;13:46-51.
Crossref - Xudong L. A Study on Environmental Pollution of Agriculture and Countermeasures under the Double Failure. Energy Procedia. 2011;5:204-208.
Crossref - Aznur BS, Nisa SK, Septriono WA. Agen Biologis Bioremediasi Logam Berat (Heavy Metal Bioremediation Biologycal Agents). MAIYAH. 2022;1(4):186.
Crossref - Tadikonda RR, Afreen D, Battula S. Green Analytical Chemistry: A Comprehensive Review of Eco-Scale, Greenness Metrics, and Sustainability Approaches. Journal of Drug Delivery and Therapeutics. 2025;15(4):179-182.
Crossref - Locatelli M, Perrucci M, de la Fuente Ballesteros A, Bernal J, Ares AM. White Analytical Chemistry: Balancing Performance, Sustainability, and Practicality in Modern Methods. LCGC International. 2025.
- Raz A, Schwartzman M, Kenig-Wakshal R. Chemical and Enzymatic Transformations of Prostaglandin Endoperoxides: Evidence for the Predominance of the 15-Hydroperoxy Pathway. Eur J Biochem. 1976;70(1):89-96.
Crossref - Reid I, Osman S, Bakheet S. Advances in Analytical Techniques for the Simultaneous Quantification of Azole Antifungals and Corticosteroids in Topical Formulations. Int J Pharm Chem. 2025;11:31-46.
Crossref - Mahajan G, Thadiyan V, Bisht T, Gupta R. Microbially Derived Chitinases: Multifunctional Enzymes for Biocontrol, Bioremediation, and Sustainable Industrial Applications. In: Arora, N.K., Pandey, P., Mishra, J. (eds) Environmental Sustainability and Biotechnology: Opportunities and Challenges. Microorganisms for Sustainability, vol 46. Springer, Singapore. 2025:165 199.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.