ISSN: 0973-7510
E-ISSN: 2581-690X
The worldwide rise of antibiotic-resistant Escherichia coli, a major cause of urinary tract infections (UTIs), has intensified concerns over patient outcomes, illness severity, and healthcare costs. This study investigated the antimicrobial resistance patterns of uropathogenic E. coli (UPEC) isolates, emphasizing the prevalence of extended-spectrum β-lactamase (ESBL) and carbapenemase producers and their associated resistance trends. A prospective study was conducted at BGS Global Institute of Medical Sciences, Bengaluru, from June 2021 to May 2022, involving 226 UPEC isolates. Antimicrobial susceptibility testing revealed high resistance to ciprofloxacin (50.9%), ampicillin (59.6%), and aztreonam (61.5%), while notable susceptibility was observed to fosfomycin (97.5%), amikacin (93.7%), and imipenem (93.4%). ESBL production was confirmed in 36.7% of isolates, and 3.1% were identified as carbapenemase producers. Molecular analysis showed that blaTEM and blaCTX genes were associated with ESBL production, while blaVIM was associated with carbapenemase production. There was no significant correlation between ESBL and carbapenemase expression. The findings highlight the alarming prevalence of multidrug-resistant UPEC and underline the necessity for continuous surveillance of antimicrobial resistance patterns. The preserved activity of fosfomycin, amikacin, and imipenem suggests their continued utility in managing resistant UTIs. Rational antibiotic use and stringent infection control measures remain imperative to curb the further spread of resistant strains.
Antibiotic Resistance, Carbapenemase Genes, ESBL (Extended-Spectrum Beta-Lactamase) Genes, Uropathogenic Escherichia coli (E. coli), Urinary Tract Infections (UTIs)
Urinary Tract Infections (UTIs) – A possible pandemic menace looming large
Among the infections of bacterial origin, Urinary tract infections (UTIs) are most prevalent and annually approximately 150 million people around the globe are affected by it.1 These infections pose a significant burden on healthcare systems, leading to high morbidity, hospitalization rates, and increased treatment costs. UTIs can affect individuals of all ages, but certain populations, such as women, diabetics, and the elderly, are more susceptible due to anatomical and immunological factors.
Pathogenic role of Escherichia coli (E. coli) in UTIs
In the reported cases of UTIs, it has been observed that a major fraction (70%-80%) is caused by uropathogenic Escherichia coli (UPEC)**,2,3 which establishes its colonies in the urinary tract epithelium and evades the host immune system using through the action of virulence factors such as: Fimbriae and adhesins (e.g., type 1 pili, P. fimbriae) – allowing bacterial attachment to urothelial cells. Toxins (e.g., hemolysin, cytotoxic necrotizing factor) – damaging host tissues and promoting bacterial survival.
UTIs can be classified into two broad categories based on epidemiology
Community-acquired UTIs – primarily caused by non-catheter-associated infections.
Nosocomial (hospital-acquired) UTIs – often associated with catheterization, prolonged hospital stays, and multidrug-resistant (MDR) pathogens.4 Understanding these classifications is essential, as the type of infection often determines the causative organisms, resistance patterns, and treatment strategies. The following section explores these aspects in greater detail, highlighting the predominant pathogens and their antimicrobial resistance profiles. Nosocomial UTIs pose a significant threat due to their association with MDR bacteria, often leading to treatment failures and prolonged hospitalizations. Reports indicate that these infections comprise about 40% of all the hospital-acquired infections and 50% of the infections of bacterial origin.5
Rising antibiotic resistance – A public health threat
The increasing resistance to antibiotics like ampicillin and ciprofloxacin among uropathogenic E. coli (UPEC) has complicated treatment strategies and led to the failure of antibiotics. The primary mechanisms of antibiotic resistance in UPEC include:
Efflux pumps
These are active transport systems that expel antibiotics and toxic compounds from bacterial cells, thereby reducing the intracellular concentration of drugs to sub-lethal levels. Efflux pumps can be constitutively expressed or induced upon exposure to antibiotics or environmental stress. Their activation often involves global regulatory genes (such as marA, soxS, and rob in E. coli) that upregulate efflux pump operons. The overexpression of these systems contributes to multidrug-resistance (MDR) by enabling bacteria to simultaneously eject multiple classes of antibiotics, including fluoroquinolones, tetracyclines, and β-lactams.
Heteroresistance
Subpopulations of bacteria display varying levels of antibiotic resistance, allowing a fraction of the population to survive antibiotic treatment and potentially develop full resistance upon further exposure.
Alteration in outer membrane permeability
Modifications in porin proteins or loss of specific channels reduce antibiotic entry into bacterial cells, thereby decreasing susceptibility to β-lactams and fluoroquinolones.
Production of β-lactamase enzymes
These enzymes hydrolyze the β-lactam ring of antibiotics such as penicillins and cephalosporins, rendering them inactive.6 The emergence of extended-spectrum β-lactamases (ESBLs) and carbapenemases has further intensified resistance in uropathogens.
Among β-lactamase enzymes, extended-spectrum β-lactamases (ESBLs) have become a major concern due to their ability to break down third-generation cephalosporins (e.g., ceftriaxone, ceftazidime, cefotaxime). Mobile genetic elements like plasmids often bear the genes that encode for ESBLs which bring about rapid horizontal gene transfer (HGT) among the bacterial populations.7
Carbapenem-Resistant Pathogens – The Emerging Crisis:
The last line of defense against ESBL producing bacteria is carbapenem antibiotics, such as imipenem, meropenem and ertapenem.8 Unfortunately, the emergence of carbapenemase- producing Enterobacteriaceae (CPE) has challenged the effectiveness of these drugs as the enzyme carbapenemase hydrolyzes carbapenems resulting in extreme drug resistance (XDR) infections. Notable carbapenemase enzymes include:
- Klebsiella pneumoniae carbapenemase (KPC)
- Verona integron-encoded metallo-β-lactamase (VIM)
- New Delhi metallo-β-lactamase (NDM)
- Oxacillinase-48 (OXA-48)
Infections caused by carbapenemase-producing bacteria have been linked to high mortality rates (up to 50%), prolonged hospital stays, and increased healthcare costs.9
Molecular Diagnostics – A Tool for Rapid Detection
Molecular diagnostic techniques, such as polymerase chain reaction (PCR), whole-genome sequencing (WGS), and multiplex PCR, have become essential tools in detecting ESBL- and carbapenemase-producing bacteria. These methods allow for:
- Early identification of resistant bacterial strains.
- Rapid differentiation of resistance genes.
- Guiding targeted antibiotic therapy to improve patient outcomes.
However, due to their high cost and requirement for advanced laboratory facilities, molecular techniques are not always accessible in resource-limited settings. Consequently, phenotypic methods like Kirby-Bauer disc diffusion test, double-disc synergy test and modified Hodge test are widely used for detection of ESBL and carbapenemase.
Given the clinical and epidemiological significance of antibiotic-resistant UPEC, this study was conducted to:
- Isolate and identify uropathogenic E. coli and study its antimicrobial resistance.
- Using phenotypic and genotypic methods to detect ESBL and carbapenemase production among UPEC isolates.
- To examine the nexus between antibiotic resistance and the production of ESBL and carbapenemase among the UPEC isolates.
This study aims to provide critical insights into antimicrobial resistance trends, which can help shape effective antibiotic stewardship programs and infection control policies to combat drug-resistant UTIs.
Patients and Methods
Study design
Upon procuring the approval from the institutional ethics committee, a prospective study was carried out from June 2021 to May 2022 at BGS Global Institute of Medical Sciences, Bengaluru. From every participating patient, a detailed informed consent was obtained and were instructed to provide mid-stream, clean-catch urine samples after proper genital toileting.
Inclusion criteria
Patients who were 18 years of age or older, exhibiting UTI symptoms, and providing their informed consent were included.
Exclusion criteria
Patients under 18 years of age, those without symptoms of UTI, and those without informed consent were excluded from the study.
Isolation and identification of UPEC isolates
Eligible patients had their urine samples taken under stringent aseptic precautions, and the samples were processed within an hour of being taken. Wet mount preparations were performed for observing pus cells, erythrocytes, bacteria, and urinary casts.
More than 5 leukocytes per high-power field were defined as significant pyuria. The presence and morphology of bacteria were examined using Gram staining method.10,11 Samples were cultured and subjected to sensitivity testing using standard protocols; and semi-quantitative urine culture analyses were performed using the calibrated loop technique, with inoculation on MacConkey agar and blood agar plates. An incubation time of 24 hours at 37 °C in aerobic conditions was considered significant when colony counts exceeded 105 CFU/mL.
Phenotypic detection
ESBLs detection
Screening method
Kirby Bauer disc diffusion test (zone of inhibition ≤17 mm) was performed using the isolates of Escherichia coli to screen for resistance to 2nd and 3rd generation cephalosporins: ceftazidime (30 µg) and cefotaxime (30 µg).12 UPEC isolates that exhibited resistance to either one or both third-generation cephalosporins, were counted as positive for ESBL production in screening test.
Confirmatory method
UPEC isolates that tested positive for ESBL production in the initial screening were further confirmed using the combination disc method. Discs containing either ceftazidime (30 µg) or ceftazidime-clavulanic acid (30 µg/10 µg), or cefotaxime (30 µg) and cefotaxime-clavulanic acid (30 µg/10 µg) were used. An increase of ≥5 mm in the zone diameter around the cephalosporin-clavulanic acid disc compared to the cephalosporin disc alone was interpreted as positive for ESBL production, according to CLSI guidelines. Escherichia coli ATCC 35218 (non-ESBL producer) and Escherichia coli ATCC 700603 (ESBL producer) were used as negative and positive controls, respectively.13
Carbapenemase detection
Screening method
UPEC isolates were screened for resistance to imipenem (10 µg) disc by Kirby-Bauer disc diffusion test.12 Isolates that displayed resistance to imipenem disc (<19 mm zone of inhibition) were considered positive for carbapenemase production in screening test.
Confirmatory method
UPEC isolates demonstrating presumptive carbapenemase production on initial screening were subjected to confirmation by the combined disc test. Imipenem (10 µg) and imipenem-EDTA (10 µg/0.5 µg) discs were placed on Mueller-Hinton agar plates inoculated with the test isolates. An increase of ≥7 mm in the inhibition zone diameter around the imipenem-EDTA disc, compared to the imipenem disc alone, was interpreted as indicative of carbapenemase production, as per standard guidelines.14-16
Anti-microbial testing
Antimicrobial susceptibility testing was conducted using the Kirby-Bauer disc diffusion method on Mueller-Hinton agar, following Clinical and Laboratory Standards Institute (CLSI) guidelines. The following antibiotic discs (HiMedia Laboratories, Mumbai, India) were employed: gentamicin (10 µg), nitrofurantoin (300 µg), cotrimoxazole (1.25/23.75 µg), norfloxacin (10 µg), aztreonam (30 µg), ceftazidime (30 µg), cefotaxime (30 µg), amikacin (30 µg), amoxicillin-clavulanic acid (30 µg), ciprofloxacin (5 µg), levofloxacin (5 µg), imipenem (10 µg), meropenem (10 µg), fosfomycin (200 µg) and piperacillin-tazobactam (100/10 µg). Zone diameters were interpreted according to CLSI breakpoints. Escherichia coli ATCC 25922 served as the quality control strain. Only isolates of E. coli exhibiting colony counts ≥105 CFU/mL were included for antimicrobial susceptibility testing and further evaluated for ESBL and carbapenemase production.13
Molecular genotyping for ESBL and Carbapenemase genes
Molecular genotyping of Escherichia coli isolates was conducted using a combination of bioinformatics tools, multiplex PCR, and colony PCR. To guide the selection of relevant resistance genes, four representative E. coli isolates were selected for Whole Genome Sequencing (WGS) using high throughput sequencing platforms. The WGS data were analyzed through bioinformatics pipelines to identify prevalent ESBL and carbapenemase genes circulating in southern India. Based on the genetic profiles derived from WGS, a targeted panel of resistance genes blaTEM, blaCTX-M, blaPER, blaIMP, blaVIM, blaNDM, blaKPC, and blaOXA-48-like was selected for screening in the remaining UPEC isolates.14,16 This WGS-guided approach enabled focused molecular detection of clinically and regionally significant antimicrobial resistance determinants.
Primer design for antibiotic resistance genes
Primer design for the targeted genes involved in ESBL and carbapenemase production was conducted using bioinformatics tools and established guidelines. The following steps outline the methodology:
Sequence retrieval
The nucleotide sequences of the target genes (blaTEM, blaCTX-M, blaPER, blaIMP, blaVIM, blaNDM, blaKPC, and blaOXA-48-like) were retrieved from databases such as GenBank or relevant literature.
Sequence Alignment and Analysis
The retrieved sequences were aligned using multiple sequence alignment tools to identify conserved regions among different variants of the target genes. Conserved regions were selected for primer design to ensure specificity and efficiency.
Primer Design Software
Bioinformatics tools such as Primer3, NCBI Primer-BLAST, or similar software were utilized for primer design. Primers were designed to flank the conserved regions identified in the sequence alignment while ensuring optimal length, GC content, melting temperature (Tm), and minimizing secondary structures.
Primer Specificity Check
The designed primers were validated in silico against the target gene sequences and other related sequences to avoid non-specific amplification. Tools such as BLAST were used to assess primer specificity.
By following these steps, specific and efficient primer pairs were designed for the detection and characterization of mechanisms of antibiotic resistance in E. coli isolates.
Bacterial DNA extraction and PCR amplification
Genomic DNA was extracted from bacterial isolates using the Genei DNA extraction kit (Genei, India), following the manufacturer’s protocol. PCR was performed in a total reaction volume of 25 µL containing 1× JumpStart REDTaq ReadyMix (Sigma-Aldrich, USA) and gene-specific primers. The thermal cycling conditions included an initial denaturation at 95 °C for 3 minutes, followed by 35 cycles of (i) Denaturation at 95 °C for 30 seconds, (ii) Annealing and extension at 62.5 °C for 1 hour 30 minutes, and (iii) A final extension at 72 °C for 5 minutes.
Negative controls lacking template DNA were included in each run to check for contamination. PCR products (5 µL) were electrophoresed on 1.5% (w/v) agarose gels prepared in 0.5 × TBE buffer and run at 100 V for 30 minutes. The gels were stained with ethidium bromide and visualized under UV transillumination. Amplicon sizes were determined by comparison with a molecular size marker.17
Bacterial colonies were cultured on LB agar for 18 hours at 37 °C. A small amount of pure colony was collected using a sterile toothpick and suspended in the PCR reaction mixture. Colony PCR reactions and gel electrophoresis were conducted using the same procedure described above.
Primers for ESBL and carbapenemase genes were designed and purchased from GENEI, and JumpStart REDTaq ReadyMix was procured from Sigma-Aldrich. Their sequences are listed in Table 1.
Table (1):
Primer Sequences for Carbapenemase and ESBL Genes
| No. | Primer | Sequence | Length | Temp. (°C) | GC% |
|---|---|---|---|---|---|
| 1 | blaTEM | GGGAACCGGAGCTGAATGAA | 20 | 60.04 | 55 |
| CAGTGCTGCAATGATACCGC | 20 | 59.97 | 55 | ||
| 2 | blaCTX-M | TTACCCAGCGTCAGATTCCG | 20 | 59.83 | 55 |
| TGCGGAAAAGCACGTCAATG | 20 | 60.04 | 50 | ||
| 3 | blaPER | GCAACTGCTGCAATACTCGG | 20 | 60.03 | 55 |
| CATCTGCGCTTCATTTG | 20 | 59.96 | 55 | ||
| 4 | blaIMP | GCTACCGCAGCAGAGTCTTT | 20 | 60.39 | 55 |
| AAAGCCCCACCCGTTAACTT | 20 | 59.81 | 50 | ||
| 5 | blaVIM | GCTACCGCAGCAGAGTCTTT | 20 | 60.39 | 55 |
| AAAGCCCCACCCGTTAACTT | 20 | 59.81 | 50 | ||
| 6 | blaNDM | CCTGGACCAATGACCAGACC | 20 | 60.04 | 60 |
| AATACCTTGAGCGGGCCAAA | 20 | 59.96 | 50 | ||
| 7 | blaKPC | TCCGTTACGGCAAAAATGCG | 20 | 60.11 | 50 |
| CGGCATAGTCATTTGCCGTG | 20 | 59.97 | 55 | ||
| 8 | blaOXA-48-like | AATTGGCGAGGCACGTATGA | 20 | 60.11 | 50 |
| TTCGAATACCACCGTCGAGC | 20 | 60.18 | 55 |
Data analysis
Data entry was done on Microsoft Excel data sheet. Data was analyzed using SPSS version [IBM SPSS Statistics, SOMERS, NY USA]. The association of ESBL and MBL with antimicrobial resistance was studied, using Chi-square test. A p-value <0.001 was considered as statistically significant.17
Characteristics of UPEC patients: A total of 226 patients were examined, of which 56.6% were females (n = 128), and 43.4% were males (n = 98) (Table 2). Most of the patients were between the ages of 18 and 25, followed by 56-65 years of age, 26-35 years of age, 36-45 years of age, 66-75 years of age, 46-55 years of age, and 76-85 years of age (Table 2). Patients with no comorbid conditions constituted the majority of the patient population (n = 116) (Table 2). A total of 105 patient isolates were found to have type-II diabetes mellitus, followed by chronic kidney disease (n = 10), and prostate carcinoma (1 patient isolate) (Table 2).
Table (2):
Demographic Distribution and Comorbidities of Patients with Urinary Tract Infection
| Age Group (Years) | Number of patients n (%) | Sex n (%) | Comorbidities | |
|---|---|---|---|---|
| Male | Female | |||
| 18-25 | 60 (26.5%) | 98 (43.4%) | 128 (56.6%) | Diabetes (105) |
| 26-35 | 28 (12.4%) | CKD (10) | ||
| 36-45 | 28 (12.4%) | Prostate carcinoma (01) | ||
| 46-55 | 21 (9.30%) | None (116) | ||
| 56-65 | 50 (22.1%) | |||
| 66-75 | 28 (12.4%) | |||
| 76-85 | 11 (4.90%) | |||
Bacterial Uropathogens
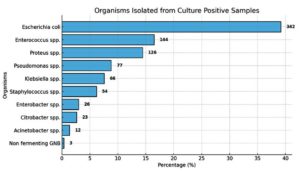
During the study period, a total of 4031 urine samples were processed, of which 873 (21.65%) yielded significant bacterial growth. Among these culture-positive samples, 226 non-repetitive isolates of uropathogenic Escherichia coli (UPEC) that met the inclusion criteria were selected for further analysis. The colony morphology of representative isolates is depicted in Figure 1. The highest proportion of UPEC isolates was recovered from patients aged 18-25 years (26.5%), followed by those aged 56-65 years (22.1%). The overall distribution of bacterial pathogens isolated from urine cultures was as follows: E. coli (39.17%), Enterococcus spp. (16.49%), Proteus spp. (14.43%), Pseudomonas spp. (8.82%), Klebsiella spp. (7.56%), Staphylococcus spp. (6.18%), Enterobacter spp. (2.97%), Citrobacter spp. (2.63%), Acinetobacter spp. (1.37%), and non-fermenting Gram-negative bacilli (0.34%) (Figure 2).
Antibiotic susceptibility pattern of E. coli
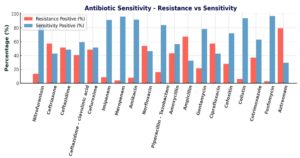
In our study, the sensitivity pattern of Escherichia coli showed good sensitivity to fosfomycin (97.5%), imipenem (93.4%), amikacin (93.7%), nitrofurantoin (88.5%), piperacillin- tazobactam (86.6%), gentamicin (82.3%), and cefoxitin (75.9%) (Figure 3). Moderate sensitivity was observed for cotrimoxazole (65.9%), ceftazidime-clavulinic acid (65.3%), ceftazidime (56.7%), ceftriaxone (52.2%), cefotaxime (54.4%), amoxicillin (63.6%), ciprofloxacin (49.1%), ampicillin (40.4%) and aztreonam (38.5%) (Figure 3).
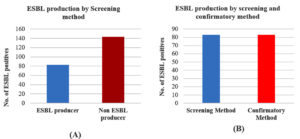
Of the 226 Escherichia coli isolates, 83 (36.7%) were identified as potential ESBL producers based on the initial screening test (Figure 4A). All isolates were subsequently subjected to phenotypic confirmation using the combined disc diffusion method. The confirmatory test also detected ESBL production in 83 (36.7%) isolates (Figure 4B). There was complete agreement between the screening and confirmatory tests, as all isolates positive by screening were confirmed as ESBL producers.
Figure 4. (A) Screening test for ESBL production among UPEC isolates; (B) ESBL production by screening method and confirmatory method
Detection of carbapenemase
In the screening method, out of 226 isolates of UPEC, 19 (8.40%) isolates were identified as carbapenemase producing UPEC isolates (Table 3). In the confirmatory method, out of 226 isolates, 7 (3.1%) isolates were identified as carbapenemase producing isolates (Table 3).
Table (3):
Carbapenemase detection by screening and confirmatory methods
Carbapenemase detection by screening method |
Number |
Percentage (%) |
|---|---|---|
UPEC isolates producing carbapenemase |
19 |
8.40 |
UPEC isolates not producing carbapenemase |
207 |
91.59 |
Carbapenemase detection by confirmatory method |
Number |
Percentage (%) |
UPEC isolates producing carbapenemase |
7 |
3.1 |
UPEC isolates not producing carbapenemase |
219 |
96.9 |
Correlation of antibiotic resistance with ESBL
Significant correlation (p < 0.005), between antibiotic resistance and ESBL production, was observed in this study for antibiotics like ceftazidime (p < 0.001), ceftazidime-clavulinic acid (p < 0.001), ceftriaxone (p < 0.001), cefoxitin (p < 0.001), norfloxacin (p < 0.001) and ciprofloxacin (p < 0.001) (Table 4).
Table (4):
Correlation of antibiotic resistance with ESBL production
Antibiotic Sensitivity |
Resistance Positive (Number) |
Resistance Positive (%) |
Resistance Negative (Number) |
Resistance Negative (%) |
Sensitivity Positive (Number) |
Sensitivity Positive (%) |
Sensitivity Negative (Number) |
Sensitivity Negative (%) |
P-value |
|---|---|---|---|---|---|---|---|---|---|
Nitrofurantoin |
24 |
13.7 |
2 |
3.9 |
151 |
86.3 |
49 |
96.1 |
0.008 |
Ceftriaxone |
100 |
57.1 |
8 |
15.7 |
75 |
42.9 |
4 |
84.3 |
0.001 |
Ceftazidime |
90 |
51.4 |
7 |
13.7 |
85 |
48.6 |
44 |
86.3 |
0.001 |
Ceftazidime-clavulinic acid |
71 |
40.6 |
4 |
7.8 |
104 |
59.4 |
47 |
92.2 |
0.001 |
Cefuroxime |
85 |
48.6 |
5 |
9.8 |
90 |
51.4 |
46 |
90.2 |
0.001 |
Imipenem |
15 |
8.6 |
0 |
0 |
160 |
91.4 |
51 |
100 |
0.09 |
Meropenem |
7 |
4 |
0 |
0 |
168 |
96 |
51 |
100 |
0.162 |
Amikacin |
14 |
8 |
0 |
0 |
161 |
92 |
51 |
100 |
0.005 |
Norfloxacin |
94 |
53.7 |
11 |
21.6 |
81 |
46.3 |
40 |
78.4 |
0.001 |
Piperacillin-Tazobactam |
28 |
16 |
2 |
3.9 |
147 |
84 |
49 |
96.1 |
0.006 |
Amoxycillin |
76 |
43.4 |
7 |
13.7 |
99 |
56.6 |
44 |
86.3 |
0.001 |
Ampicillin |
118 |
67.4 |
16 |
31.4 |
57 |
32.6 |
35 |
68.6 |
0.001 |
Gentamycin |
38 |
21.7 |
2 |
3.9 |
137 |
78.3 |
49 |
96.1 |
0.001 |
Ciprofloxacin |
100 |
57.1 |
14 |
27.5 |
75 |
42.9 |
37 |
72.5 |
0.001 |
Cefoxitin |
49 |
28 |
3 |
5.9 |
126 |
72 |
48 |
94.1 |
0.001 |
Colistin |
11 |
6.3 |
0 |
0 |
164 |
93.7 |
51 |
100 |
0.056 |
Cotrimoxazole |
65 |
37.1 |
8 |
15.7 |
110 |
62.9 |
43 |
84.3 |
0.001 |
Fosfomycin |
5 |
2.9 |
0 |
0 |
170 |
97.1 |
51 |
100 |
0.275 |
Aztreonam |
139 |
79.43 |
0 |
0 |
36 |
29.57 |
51 |
100 |
0.001 |
Genotypic expression among UPEC isolates
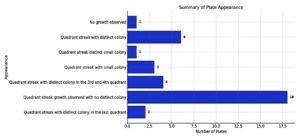
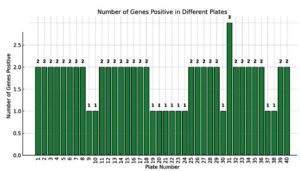
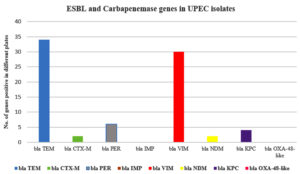
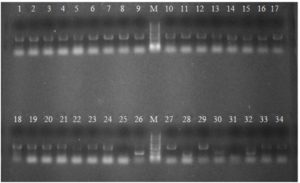
Approximately 40 genes were identified in Escherichia coli (E. coli) isolates (Figures 5 and 6). Carbapenemase genes were detected in 39 E. coli isolates. The most prevalent gene was blaTEM, present in all plates except one. Other commonly detected genes included blaVIM, found in all plates, and blaCTX, which was present in all but two plates (Figure 7).
Genes blaNDM, blaKPC, blaOXA, and blaPER were not detected in any of the bacterial isolates (Figure 5 and Table 5).
Table (5):
Distribution of ESBL and carbapenemase genes among UPEC isolates
ESBL/Carbapenemase genes |
Frequency |
% |
|---|---|---|
blaTEM |
34 |
85 |
blaCTX-M |
2 |
5 |
blaPE R |
6 |
15 |
blaIMP |
0 |
0 |
blaVIM |
30 |
75 |
blaNDM |
2 |
5 |
blaKPC |
4 |
10 |
blaOXA-48-like |
0 |
0 |
Figure 6. Average number of genes expressed in E. coli isolates linked to ESBL and Carbapenemase production
This study provides a detailed analysis of 226 non-repetitive uropathogenic Escherichia coli (UPEC) isolates, combining phenotypic and genotypic characterization of antimicrobial resistance due to ESBL and carbapenemase production. Uropathogenic E. coli (UPEC) constitutes the principal etiological agent of UTI. It has an arsenal of virulence determinants and can acquire and disseminate antibiotic resistance genes.18,19 The findings highlight the alarming emergence of multidrug-resistance and reveal the interplay between virulence and resistance determinants in urinary pathogens.
Out of 4031 urine samples, 873 (21.65%) showed significant bacterial growth, of which E. coli constituted 39.17%, confirming it as the predominant urinary pathogen. These results echo the classic observations of Ronald,3 and Foxman,20 who established E. coli as the leading cause of both community- and hospital-acquired UTIs. The higher infection rate in females (56.6%) compared to males (43.4%) corresponds with the findings of Dielubanza & Schaeffer19 and Sobel & Kaye,21 attributing this pattern to anatomical factors such as shorter urethra and hormonal predisposition.
The 18-25 year group accounted for 26.5% of the cases, followed by the 56-65 year group (22.1%), a bimodal distribution typical for sexually active females and postmenopausal women. Among comorbidities, Type-II diabetes mellitus (46%) dominated, similar to Niranjan and Kabugo et al, 22,23 who linked diabetes with impaired host defenses and recurrent UTI risk. The antibiotic sensitivity profile revealed a dual scenario – a few highly effective agents contrasted against widespread resistance to conventional antibiotics. In our cohort, E. coli isolates exhibited excellent sensitivity to fosfomycin (97.5%), amikacin (93.7%), imipenem (93.4%), and nitrofurantoin (88.5%). This aligns with Sabharwal & Sharma24 and Sultan et al.,25 who highlighted fosfomycin’s renewed therapeutic role against multidrug-resistant urinary pathogens. Conversely, resistance was markedly high to ampicillin (67.4%), ceftriaxone (57.1%), ceftazidime (51.4%), ciprofloxacin (57.1%), and aztreonam (79.4%). Comparable resistance profiles were documented by Mukherjee et al.26 in Indian tertiary centers, where empirical use of β-lactams and fluoroquinolones facilitated plasmid-borne resistance spread.
The current data further demonstrate a statistically significant (p < 0.001) correlation between ESBL production and resistance to key β-lactams and fluoroquinolones – including ceftazidime (51.4% resistant, p = 0.001), ceftriaxone (57.1% resistant, p = 0.001), cefoxitin (28%, p = 0.001), and ciprofloxacin (57.1%, p = 0.001). This co-resistance supports the assertion by Adamus-Bialek et al.2 that resistance in E. coli is frequently driven by co-located resistance genes on mobile plasmids, conferring multidrug-resistance and adaptive survival advantage.
ESBL production and molecular determinants of the 226 isolates, 83 (36.7%) were confirmed as ESBL producers by both screening and confirmatory assays. This prevalence is consistent with Bajpai et al.,17 who reported ESBL rates ranging from 30%-45% among urinary E. coli isolates in Indian hospitals.
Genotypically, the study detected blaTEM and blaCTX-M as the most dominant genes, both present in nearly all ESBL-producing isolates, while blaVIM was detected in several isolates associated with carbapenemase activity. Genes such as blaNDM, blaKPC, and blaOXA were absent.
The predominance of blaTEM agrees with Bajpai et al.,17 who identified TEM as the principal β-lactamase gene in Indian E. coli isolates, followed by blaCTX-M.26 The co-occurrence of blaTEM and blaCTX-M signifies a progressive genetic transition from classical TEM/SHV enzymes to CTX-M-type ESBLs, consistent with the molecular epidemiology described by Adamus-Bialek and Agarwal et al.2,4
The ESBL positivity was also significantly associated with multidrug-resistance patterns – isolates expressing ESBLs showed 2-4 times higher resistance to fluoroquinolones and aminoglycosides compared to non-ESBL strains, corroborating Ranjini et al and Sajama & Premlatha.13,27 Such co-selection likely reflects plasmid-mediated horizontal gene transfer encompassing blaCTX-M, aac(6’)-Ib-cr, and qnr determinants, as noted in global genomic studies.7,28
Carbapenemase detection revealed 19 isolates (8.4%) positive by screening but only 7 isolates (3.1%) confirmed phenotypically. This finding indicates possible overestimation by screening assays, a discrepancy similar to that reported by Haji et al.16 and Vamsi et al.15 The limited occurrence of carbapenemase producers in our cohort is encouraging compared to the 6%-10% prevalence documented by Mariappan et al.9 in Indian intensive care settings.
The genetic screening identified blaVIM as the only carbapenemase gene present, consistent with Legese et al.8 who reported dominance of VIM-type metallo-β-lactamases in African E. coli isolates. The absence of blaNDM and blaKPC contrasts with northern Indian and Middle Eastern reports29,30 Rahdar et al., El Naggar et al., suggesting regional containment or restricted plasmid dissemination in our hospital ecosystem.
Clinically, the carbapenem susceptibility remained high (imipenem 93.4%, meropenem 96%), supporting the continued efficacy of carbapenems in our setting.31-33 However, as emphasized by Bloemberg et al.,14 the presence of even a few carbapenemase producers represents a sentinel warning for potential clonal outbreaks, warranting strict antibiotic stewardship and infection-control protocols.34
When compared with other regional and international datasets, our 36.7% ESBL and 3.1% carbapenemase rates are lower than the figures from North India (ESBL 52%, carbapenemase 7%) but closely resemble European and Latin American averages (30%-40%).7,35 The molecular profile – blaTEM and blaCTX-M without blaNDM – suggests that plasmid-mediated dissemination remains the primary resistance mechanism in this locale, as opposed to transposon-mediated metallo-β-lactamase spread.
Therapeutically, the study supports fosfomycin, nitrofurantoin, and amikacin as viable empirical options for uncomplicated UTIs, consistent with CLSI guidelines (2015). For complicated or catheter-associated UTIs, carbapenems remain effective but must be preserved through stringent antimicrobial stewardship.
From a public-health standpoint, the coexistence of blaTEM, blaCTX-M, and carbapenem tolerance presents a significant nosocomial hazard. The findings underscore the urgency of integrating molecular diagnostics into infection surveillance frameworks.
Despite its comprehensive phenotypic–genotypic design, this study was limited by its single-center scope and lack of advanced molecular typing (e.g., whole-genome sequencing or MLST). Expanding to multicentric studies incorporating plasmid profiling and virulome-resistome linkage would strengthen the understanding of UPEC pathoadaptation.
Accurate detection of extended-spectrum β-lactamase (ESBL) and carbapenemase production among E. coli isolates causing urinary tract infections (UTIs) is pivotal for optimizing antimicrobial therapy and minimizing treatment failures. The rising prevalence of multidrug-resistant (MDR) uropathogenic E. coli (UPEC) strains underscores the need for early identification of resistance mechanisms to reduce morbidity and prevent nosocomial dissemination of resistant pathogens. The high concordance observed between phenotypic and genotypic detection methods in this study reinforces the importance of an integrated diagnostic approach. While phenotypic assays such as disc diffusion and carbapenemase detection remain practical and cost-effective for routine laboratory screening, molecular techniques like PCR-based identification of blaTEM, blaVIM, and blaKPC genes offer superior specificity and precision. The combined use of these complementary methods enhances diagnostic accuracy, thereby supporting evidence-based antimicrobial stewardship and informed therapeutic decision-making.The emergence of ESBL and carbapenemase-producing UPEC isolates in tertiary care settings presents a significant threat to public health by limiting the efficacy of β-lactam antibiotics and increasing dependence on last-line agents. These findings emphasize the urgent need for comprehensive infection control measures, periodic antimicrobial resistance surveillance, and the development of targeted antibiotic policies to preserve treatment efficacy. In the broader context of combating antimicrobial resistance, early detection, continuous monitoring, and strengthened interdisciplinary collaboration among clinicians, microbiologists, infection control personnel, and policymakers are essential for guiding empirical therapy, refining hospital antibiotic protocols, and ensuring sustainable antibiotic utilization within healthcare systems.
ACKNOWLEDGMENTS
The authors would like to thank the Management and Dean for their support. The authors also thank the teaching and non-teaching staff of the Department of Microbiology for their support, and special thanks to Mr. Aravind G, Managing Director, Dextrose Technologies Private Limited, for guiding us in the molecular run of this study.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
Both authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the Institutional Ethical Committee, BGS Global Institute of Medical Sciences, Bengaluru vide protocol number sBGS GIMS/IEC/ dated 01.02.2021.
INFORMED CONSENT
Written informed consent was obtained from the participants before enrolling in the study.
- Bergsten G, Wullt B, Svanborg C. Escherichia coli, fimbriae, bacterial persistence and host response induction in the human urinary tract. Int J Med Microbiol. 2005;295(6-7):487-502.
Crossref - Adamus-Bialek W, Baraniak A, Wawszczak M, et al. The genetic background of antibiotic resistance among clinical uropathogenic Escherichia coli strains. Mol Biol Rep. 2018;45(5):1055-1065.
Crossref - Ronald A. The etiology of urinary tract infection: traditional and emerging pathogens. Am J Med. 2002;113(Suppl 1A):14-19.
Crossref - Agarwal J, Srivastava S, Singh M. Pathogenomics of uropathogenic Escherichia coli. Ind J Med Microbiol. 2012;30(2):141-149.
Crossref - Washington CW Jr, Stephen DA, William M Janda, et al. Koneman’s colour atlas and textbook of diagnostic microbiology. 7th ed. Philadelphia: Lippincott Williams & Wilkins. 2017. Chapter 17, 1074-1121, 1127-1146.
- Stephanie Jacobson, MS MCS (ASCP)CM. Bailey & Scott’s Diagnostic Microbiology Fourteenth Edition, Elsevier US/UK 2016 Section 7, Chapter 19;329-54.
- Kazemian H, Heidari H, Ghanavati R, et al. Phenotypic and Genotypic Characterization of ESBL-, AmpC-, and Carbapenemase-Producing Klebsiella pneumoniae and Escherichia coli Isolates. Med Princ Pract. 2019;28(6):547-551.
Crossref - Legese MH, Weldearegay GM, Asrat D. Extended-spectrum beta-lactamase- and carbapenemase-producing Enterobacteriaceae among Ethiopian children. Infect Drug Resist. 2017;10:27-34.
Crossref - Mariappan S, Sekar U, Kamalanathan A. Carbapenemase-producing Enterobacteriaceae: Risk factors for infection and impact of resistance on outcomes. Int J Appl Basic Med Res. 2017;7(1):32-39.
Crossref - WWC Topley, Graham S, Sir Wilson Topley, et al. Topley and Wilson’s Microbiology and Microbial Infections 10th Edn. Distributed in the United States of America by ASM Press, London: Hodder Arnold;Washington, D.C549617.
- Winn W, Allen S, Janda W, Koneman E, et al. Koneman’s Color Atlas and Textbook of Diagnostic Microbiology 6th Edn. Philadelphia: Lippincott, New York. 2006;1443-535.
- Clinical and Laboratory Standards Institute (CLSI). M-100-S25. Performance standards for antimicrobial susceptibility testing CLSI approved standard. Clinical and Laboratory Standards Institute, Wayne. 2015.
- Ranjini C, Kasukurthi LR, Madhumati B, Rajendran R. Prevalence of multidrug resistance and extended spectrum beta-lactamases among uropathogenic Escherichia coli isolates in a tertiary care hospital in South India: An alarming trend. Community Acquired Infect. 2015;2(1):19-24.
Crossref - Bloemberg GV, Braun-Kiewnick A, Tedrup J, et al. Evaluation of AID carbapenemase line probe assay for rapid detection and identification of carbapenemase genes in Gram-negative bacilli. J Antimicrob Chemother. 2017;72(7):1948-1954.
Crossref - Vamsi SK, Moorthy RS, Hemiliamma MN, Reddy RBC, Chanderakant DJ, Sirikonda S. Phenotypic and genotypic detection of carbapenemase production among gram-negative bacteria isolated from hospital-acquired infections. Saudi Med J. 2022;43(3):236-243.
Crossref - Haji SH, Aka STH, Ali FA. Prevalence and characterisation of carbapenemase encoding genes in multidrug-resistant Gram-negative bacilli. PLoS One. 2021;16(11):e0259005.
Crossref - Bajpai T, Pandey M, Varma M, Bhatambare GS. Prevalence of TEM, SHV, and CTX-M Beta-Lactamase genes in the urinary isolates of a tertiary care hospital. Avicenna J Med. 2017;7(1):12-16.
Crossref - Nandagopal B, Sankar S, Sagadevan K, et al. Frequency of extended spectrum β-lactamase-producing urinary isolates of Gram-negative bacilli among patients seen in a multispecialty hospital in Vellore district, India. Indian J Med Microbiol. 2015;33(2):282-285.
Crossref - Dielubanza EJ, Schaeffer AJ. Urinary tract infections in women. Medical Clinics of North America. 2011;95(1):27-41.
Crossref - Foxman B. Epidemiology of urinary tract infections: incidence, morbidity, and economic costs. Am J Med. 2002;113(Suppl 1A):5S-13S.
Crossref - Sobel JD, Kaye D. Urinary tract infections. In: Mandell GL, Bennett JE, Dolin R, eds. Principles and Practice of Infectious Diseases.6th ed. Philadelphia, PA: Elsevier Churchill Livingstone; 2005:662-690
- Niranjan V, Malini A. Antimicrobial resistance pattern in Escherichia coli causing urinary tract infection among inpatients. Indian J Med Res. 2014;139(6):945-948
- Kabugo D, Kizito S, Ashok DD, et al. Factors associated with community-acquired urinary tract infections among adults attending an assessment center, Mulago Hospital Uganda. Afr Health Sci. 2016;16(4):1131-1142.
Crossref - Sabharwal ER, Sharma R. Fosfomycin: an alternative therapy for the treatment of UTI amidst escalating antimicrobial resistance. J Clin Diagn Res. 2015;9(12):DC06-DC09.
Crossref - Sultan A, Rizvi M, Khan F, Sami H, Shukla I, Khan HM. Increasing antimicrobial resistance among uropathogens: Is fosfomycin the answer? Urol Ann. 2015;7(1):26-30.
Crossref - Mukherjee M, Basu S, Mukherjee SK, Majumder M. Multidrug-resistance and extended-spectrum beta-lactamase production in uropathogenic E. coli isolated from hospitalized patients in Kolkata, India. J Clin Diagn Res. 2013;7(3):449-453.
Crossref - Sajjanar V, Premlatha DE. Detection of biofilm formation, extended spectrum beta lactamase production and their correlation with antibiotic resistance among uropathogenic Escherichia coli. IP Int J Med Microbiol Trop Dis. 2021;7(3):141-147.
Crossref - Whelan S, Bottacini F, Buttimer C, Finn K, Lucey B. Whole genome sequencing of uropathogenic E. coli from Ireland reveals diverse resistance mechanisms and strong correlation with phenotypic (EUCAST) susceptibility testing. Infect Genet Evol. 2024;121:105600.
Crossref - Rahdar HA, Malekabad ES, Dadashi AR, et al. Correlation between carbapenem resistance and antibiotic resistance among clinical isolates of Klebsiella pneumoniae. Ethiop J Health Sci. 2019;29(6):745-750.
Crossref - El Naggar NM, Shawky RM, Serry FME, Emara M. Investigating the relationship between carbapenemase production and biofilm formation in Klebsiella pneumoniae clinical isolates. BMC Res Notes 2024;17(1):49.
Crossref - Kashid RA, Joseph S, Hamdan U. Antibiotic resistance pattern of uropathogens, in a tertiary care hospital in South India. CIBTech Journal of Microbiology. 2019;8(2):10-22
- Mahesh E, Ramesh D, Indumathi VA, et al. Risk factors for community-acquired urinary tract infection caused by ESBL-producing bacteria. JIACM. 2010;11(4):271.
- Ponnusamy P, Nagappan R. Extended spectrum beta-lactamase, biofilm-producing uropathogenic pathogens and their antibiotic susceptibility patterns from urinary tract infection-an overview. Int J Microbiol Res. 2013;4(2):1836-842.
- Somashekara SC, Deepalaxmi S, Jagannath N, Ramesh B, Laveesh MR, Govindadas D. Retrospective analysis of antibiotic resistance pattern to urinary pathogens in a Tertiary Care Hospital in South India. J Basic Clin Pharm. 2014;5(4):105-108.
Crossref - Ballesteros-Monrreal MG, Mendez-Pfeiffer P, Ortiz B, et al. Uropathogenic E. coli and Hybrid Pathotypes in Mexican Women with Urinary Tract Infections: A Comprehensive Molecular and Phenotypic Overview. Curr Issues Mol Biol. 2024;46(6):5909-5928.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.