ISSN: 0973-7510
E-ISSN: 2581-690X
Bloodstream infections are associated with high morbidity and mortality. Blood culture remains the gold standard for diagnosis. Inadequate incubation of the solid media subculture plates may result in delayed diagnosis. This study aimed to evaluate the diagnostic yield of extended incubation of subculture plates from conventional liquid blood cultures, analyse the recovered isolates, and determine the rate of contamination during extended incubation. This prospective, laboratory-based observational study was conducted in the Department of Microbiology at a tertiary care teaching hospital from January to March 2025. A total of 851 one-day-old sterile subculture plates were further incubated at 37 °C for another five days (i.e., day 2 to day 6) and observed daily for growth of potential pathogenic bacteria or visible contamination. Plates showing visible contamination were discarded, while those without contamination were further processed for identification of potential bacterial pathogens using routine laboratory procedures. Extended incubation yielded 29 additional potential pathogens (3.4%), predominantly Enterococcus spp. (41.5%), Acinetobacter spp. (20.7%), Staphylococci other than Staphylococcus aureus (20.7%) and Candida tropicalis (6.9%). Plate contamination rates after six days were 13.4% for blood agar and 9.0% for MacConkey agar. Contamination remained within acceptable limit (≤5%) after three days of incubation for both agar and after four days for only MacConkey agar, indicating a safe diagnostic window for extended incubation. Prolonging incubation can improve the detection of slow-growing pathogens with minimal contamination risk, even after the third or fourth day of incubation. This low-cost strategy is particularly relevant for resource-limited settings and could prevent missed diagnoses.
Bloodstream Infection, Extended Incubation, Blood Culture, Slow-growing Bacteria, Contamination Rate
Bloodstream infections (BSI) occur when bacteria, fungi, or viruses gain access to the bloodstream. They are severe clinical conditions associated with high morbidity and mortality. BSI frequently lead to sepsis, septic shock, and multi-organ dysfunction if not promptly detected and treated.1,2 According to the World Health Organization, data published in 2020 estimated that approximately 48.9 million cases of sepsis and 11 million sepsis-related deaths occurred globally, accounting for nearly one-fifth of all worldwide deaths. In hospital settings, sepsis affects an estimated 15 per 1,000 hospitalized patients, with disproportionately higher incidence and mortality observed in low and middle-income countries. In addition to its substantial clinical burden, sepsis imposes significant economic costs, with average per-patient hospital expenditures exceeding US$ 32,000 in high-income settings.3,4 The rising problem of antimicrobial resistance (AMR) further complicates the situation by reducing therapeutic options. The pathogens implicated in BSI may vary based on geography and patient population. Most common organisms worldwide include Escherichia coli, Staphylococcus aureus, Klebsiella pneumoniae, Candida species, and Staphylococci other than Staphylococcus aureus.5,6 According to Indian reports, there is a sharp rise in the prevalence of multidrug-resistant (MDR) organisms like methicillin-resistant S. aureus (MRSA) and vancomycin-resistant Enterococcus species.7
Early and accurate detection of BSI is the cornerstone of patient management, as delayed or missed detection is associated with poor outcomes.8 Blood culture remains the gold standard for BSI diagnosis, though associated with limitations like delayed positivity and potential false negatives.9 Although rapid diagnostic methods for pathogen identification and AST have been developed and evaluated to facilitate prompt clinical decision making,10 diagnostic speed alone does not guarantee optimal patient care. Some clinically significant pathogens, particularly slow-growing and fastidious organisms, may require prolonged incubation for reliable detection. Extending the incubation of blood culture samples that are initially negative, especially when slow-growing organisms are present, may improve diagnostic yield and ultimately save time by avoiding the need for repeat blood culture testing due to missed diagnoses.
The standard incubation time for blood culture varies from five to seven days for both automated and conventional blood culture methods.11 Conventional or manual blood culture may be the only option available at many peripheral, resource-limited settings due to its significantly lower cost.12 Conventional blood culture workflows involve subculturing positive bottles onto solid media such as blood agar and MacConkey agar to obtain isolated colonies for organism identification and antimicrobial susceptibility testing. These solid media are routinely incubated for only 24-48 hrs, which may be insufficient for the recovery of fastidious or slow-growing organisms. Previous studies have primarily focused on optimizing liquid blood culture systems, demonstrating that an incubation period of 4-7 days is sufficient for reliable detection of most clinically significant pathogens.13-15 While these studies clearly establish evidence for optimal incubation duration in liquid culture media, they do not address whether similar principles apply to solid media subcultures derived from positive blood culture broths. Consequently, data evaluating the impact of extended incubation of solid media on organism recovery remains scarce. This gap in evidence, particularly for fastidious or slow-growing bacteria, provides the rationale for the present study, which evaluates whether extending incubation of subculture plates beyond the conventional period improves recovery of slow-growing organisms in suspected bloodstream infections.
Study setting
This prospective laboratory-based observational study was carried out in the Department of Microbiology at the Institute of Medical Sciences, Banaras Hindu University, Varanasi, Uttar Pradesh, India, from January 2025 to March 2025.
Ethical
The study was conducted prospectively after obtaining approval from the Institutional Ethics Committee (IEC Approval No. Dean/2024/EC/7014, dated 28.02.2024).
Sample selection criteria and sample collection
All eligible, non-duplicate subculture plate sets were prospectively collected during the study period from the routine diagnostic bacteriology laboratory, Department of Microbiology, Institute of Medical Sciences, Banaras Hindu University. Each study sample consisted of one 5% blood agar (BA) and one MacConkey agar prepared from conventional liquid blood culture broths that had been reported sterile after overnight (one day) aerobic incubation at 37 °C. Plate sets incubated only for one day and showing no growth were included. Plate sets incubated for more than one day, showing mixed bacterial growth on extended incubation, or lacking patient details were excluded.
Extended incubation procedure
The collected one-day-old subculture plate sets were further incubated at 37 °C for another five days (i.e., day 2 to day 6) and observed daily for growth of potential pathogenic bacteria or visible contamination. Plates showing visible contamination were discarded, while those without contamination were further processed for identification of potential bacterial pathogens using routine laboratory procedures including Gram staining (HiMedia Laboratories, India), conventional biochemical tests (HiMedia Laboratories, India) and Vitek-2 (bioMérieux, France).
Contamination rate calculation
The rate of plate contamination was calculated using the formula:
Contamination Rate (%) = Number of contaminated plates / Total number of plates processed × 100
Quality Control (QC)
Quality control was ensured throughout the study in accordance with Clinical and Laboratory Standards Institute (CLSI) recommendations. Standard ATCC strains were used for routine internal quality checks of culture media and biochemical tests. Media sterility and performance were verified before use by incubation of uninoculated plates and by assessing the growth of reference strains. The laboratory also participates in external quality assurance scheme (EQAS) programs regularly to ensure accuracy and reliability of results.
Data collection and analysis
Demographic data, including age and gender, were obtained from patient test requisition forms. The data were entered into Microsoft Excel, which was also used for analysis. Descriptive statistics were applied, and the results were expressed as percentages. Graphical representation of data was performed using pie charts and bar diagrams.
Gender and age distribution
Among all received specimens during the study period, the majority (57.2%) were from male patients. However, among the specimens showing pathogen growth on extended incubation, the gender distribution was nearly equal, with males accounting for 51.7% and females for 48.3% (Table). The majority of patients overall were over 45 years of age (33%), followed by the 1-15 years age group (28.9%), with the median age being 28 years (standard deviation 24.9 years). However, among the specimens showing pathogen growth on extended incubation, the largest portion (31%) was formed by the 16-30 year age group, with a median age of 24 years and a standard deviation of 23.2 years, respectively (Table).
Table:
Demographic characteristics of patients and distribution of bacterial isolates recovered on extended incubation
| Variables | Overall included specimens n (%) N = 851 | Specimens with growth on extended incubation n (%) N = 29 | Bacterial species isolated*n (% within each species) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Enterococcus spp. (N = 12) | Acinetobacter spp. (N = 6) | SOSA (N = 6) | Candida tropicalis (N = 2) | S. aureus (N = 1) | K. pneumoniae (N = 1) | E. coli (N = 1) | ||||
| Sex | Male | 487 (57.2%) | 15 (51.7%) | 7 (58.3%) | 3 (50%) | 3 (50.0%) | 1 (50.0%) | – | 1 (100%) | – |
| Female | 364 (42.8%) | 14 (48.3%) | 5 (41.7%) | 3 (50%) | 3 (50.0%) | 1 (50.0%) | 1 (100%) | – | 1 (100%) | |
| Age group (year) | <1 | 48 (5.6%) | 2 (6.9%) | – | 1 (16.7%) | – | 1 (50.0%) | – | – | – |
| 1-15 | 246 (28.9%) | 7 (24.1%) | 3 (25%) | 1 (16.7%) | 1 (16.7%) | 1 (50.0%) | 1 (100%) | – | – | |
| 16-30 | 166 (19.5%) | 9 (31.0%) | 5 (41.7%) | 3 (49.9%) | 1 (16.7%) | – | – | – | – | |
| 31-45 | 111 (13.0%) | 3 (10.3%) | 1 (8.3%) | 1 (16.7%) | 1 (16.7%) | – | – | – | – | |
| >45 | 280 (33.0%) | 8 (27.6%) | 3 (25%) | – | 3 (49.9%) | – | – | 1 (100%) | 1 (100%) | |
Values are presented as n (%). Extended incubation refers to incubation of solid subculture media beyond the conventional 24-48 hrs. N denotes the total number of specimens or isolates in each category. “-” indicates no isolates recovered.
Abbreviations: SOSA: Staphylococci other than Staphylococcus aureus; spp: species; S. aureus: Staphylococcus aureus; K. pneumoniae: Klebsiella pneumoniae; E. coli: Escherichia coli
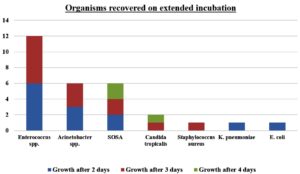
Organisms recovered on extended incubation
Out of the total 851 samples, organisms were recovered in 29 as a result of extended incubation (3.4%). Majority of these organisms were Enterococcus species (12, 41.5%), followed by Acinetobacter species (6, 20.7%), Staphylococci other than Staphylococcus aureus (6, 20.7%), Candida tropicalis (2, 6.9%), Staphylococcus aureus (1, 3.4%), Klebsiella pneumoniae (1, 3.4%), and Escherichia coli (1, 3.4%). The sex wise and age group wise distribution of these isolates is summarized in Table. Out of these six Enterococcus species, three Acinetobacter species, two Staphylococci other than Staphylococcus aureus (SOSA), one Klebsiella pneumoniae, and one Escherichia coli grew after two days of aerobic incubation. Six Enterococcus species, three Acinetobacter species, two Staphylococci other than Staphylococcus aureus, one Candida tropicalis, and one Staphylococcus aureus grew after three days of aerobic incubation. One Candida tropicalis and one Staphylococcus other than Staphylococcus aureus grew after four days of aerobic incubation (Figure 1).
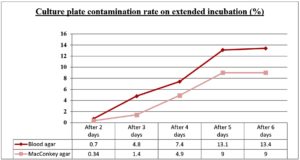
Rate of plate contamination on extended incubation
Blood agar and MacConkey agar. On Blood agar, contamination increased from 0.7% after two days to 4.8% after three days, and 7.4% after four days, followed by a sharp rise to 13.1% after five days and 13.4% after six days. MacConkey agar showed lower rates, rising from 0.34% after two days to 1.4% after three days and 4.9% after four days, then increasing to 9% after five days, with no further change by day six. Across all time points, Blood agar consistently had higher contamination rates than MacConkey agar (Figure 2).
Bloodstream infections (BSI) may range from self-limiting mild infections to sepsis with high morbidity and mortality, particularly among hospitalized patients. The importance of timely diagnosis of bloodstream infections can be appreciated from the global burden of sepsis, for which BSIs are a major underlying cause. According to the World Health Organization, sepsis affects nearly 49 million individuals annually and accounts for approximately 11 million deaths worldwide, highlighting the scale of severe infections arising from bloodstream involvement. Delayed or missed diagnosis of BSIs contributes to inappropriate or delayed targeted antimicrobial therapy, leading to increased morbidity, mortality, prolonged hospitalization, and substantial healthcare costs.4
In the present study, 57.2% of the total specimens and 51.7% of the specimens that yielded pathogens on extended incubation were from male patients. Although no statistically significant association between gender and the type of isolated organisms was observed, the predominance of males in our cohort is consistent with findings reported by previous studies.16-18 Dias et al. justified that a higher report of infections in males could be due to stronger immune responses in females, better health-seeking behaviour, and quality of health care in males.19 This could explain the findings of our study.
Majority (33%) of the total specimens received in the present study were from the elderly age group (>45 years), whereas among the specimens that yielded pathogens on extended incubation majority were from the 16-30 years age group (31%), followed by the elderly age group (27.6%). These findings are in concordance with those of other authors. Uslan et al. concluded a sharp rise in BSI incidence with increasing age.17 In a study at a tertiary hospital in western India, Sharma et al. observed majority of the BSI infection patients belonged to extreme age groups, which was attributed to age-associated immunodeficiency.18 A high positivity of blood culture has also been observed in the age group 11-30 years in another study from the Rajasthan region, wherein over 30% of the patients were in this age group.20 Possible explanations for the high positivity in this group could be behavioural and occupational exposure, as well as differences in access to and utilization of healthcare services. Collectively, these results emphasize the diagnostic value of extended incubation methods across genders and a broad range of age groups, and their applicability should not be restricted to particular demographic groups.
The optimal duration of blood culture incubation has been a well-researched topic in clinical microbiology; however, most investigations have focused on liquid blood culture systems, and there is a paucity of data directly evaluating or comparing incubation durations for solid media subcultures. Although many studies support shorter incubation periods of up to 5-7 days for liquid blood culture broths to facilitate earlier detection and reduce contamination, some also continue to advocate extended incubation durations of more than 7 days to ensure recovery of fastidious organisms, slow-growing bacteria and fungi, as well as phenotypically slow-growing variants of typically non-fastidious pathogens.2,13-15 This consideration is particularly relevant for the manual blood culture method, which remains widely used in developing countries. Yagupsky in their review, concluded that most Brucella species grow within seven days in automated blood culture systems, but growth is slower in manual blood culture media like Castaneda, which may require incubation up to two to three weeks.13 Ransom et al., in their study involving 13,592 positive blood cultures and simulated HACEK specimens, observed that there was no added benefit or increased positivity of blood culture bottles after four days of incubation.14 Nonetheless, optimum incubation remains critical to ensure adequate growth, isolation, and subsequent identification of pathogens causing bloodstream infections.
In the present study, we evaluated the extended incubation (following routine one-day incubation) of the blind subcultures done from the manual blood culture broths and its impact on improved recovery of potentially pathogenic organisms. An increased recovery rate of 3.4% was observed (29/851). The majority among them were Enterococcus species (41.5%). Other organisms isolated were Acinetobacter species (20.7%), Staphylococci other than Staphylococcus aureus (20.7%), Candida tropicalis (6.9%), Staphylococcus aureus (3.4%), Klebsiella pneumoniae (3.4%), and Escherichia coli (3.4%). Candida species, including Candida tropicalis colonies, normally start growing after two to three days, and may remain undetected if the subcultures are not routinely incubated for this period.21 Various factors that can delay or prolong the growth of microorganisms include low initial inoculum, temperature fluctuations, culture media quality, and environmental stressors such as prior antibiotic exposure. Most of the patients admitted to tertiary hospitals like ours have a history of exposure to multiple antibiotics. He and Gallert concluded in their study that sub-inhibitory concentrations of antibiotics may reduce the growth rate of organisms, including Enterococcus, E. coli and Staphylococcus species.22
One of the major concerns associated with extended incubation, especially on solid culture media plates, is contamination. The rate of contamination increases proportionally with the number of incubation days. It was observed in the present study that 13.4% of the blood agar and 9% of the MacConkey agar plates were contaminated after six days of aerobic incubation. Common plate contaminants observed included aerobic spore bearers (ASB), fungi, diphtheroids, and micrococci. Bacterial contamination was more common than fungal contamination, likely due to the longer generation time of fungi, whereas surface bacterial contaminants are typically detectable within one to two days of inoculation.23 Several factors may contribute to plate contamination, including a contaminated work environment, incubator-associated contamination, and the deposition of water droplets from condensation onto the agar surface. In incubators equipped with fans, spores of contaminant bacteria and fungi can be disseminated and enter the plates through the gap between the Petri dish and lid. Frequent handling and inspection of culture plates, especially outside a laminar flow hood, further increase the risk of airborne contamination.23-25 In the present study, this was likely a major contributor, as plates were observed at least once daily for growth. Identifying plate contamination is crucial. Indicators include growth near the plate edges, colonies forming away from inoculation streak lines, or the presence of large mould-like colonies usually indicates contamination.26,27 Furthermore, we observed a higher rate of contamination in Blood agar than in MacConkey agar. This may be due to the more enriched nature of Blood agar and the mild inhibitory nature of MacConkey agar towards Gram-positive organisms. In the absence of a standard reference for an acceptable contamination rate on general bacteriology solid media, the CDC-recommended threshold of ≤5% for mycobacterial cultures on solid media was adopted as a comparative benchmark for our study.28 Contamination was within an acceptable limit for both agars after the second and third day, and for MacConkey agar even after four days of incubation, indicating incubation could be safely extended up to two to three days. Nonetheless, plate contamination should be minimised to ensure reliable results. Measures include proper media preparation, thorough sterilization of glassware, use of freshly prepared media, and avoidance of reusing streaked sterile plates. Strict aseptic practices such as hand hygiene, use of gloves and masks, regular cleaning of the work surfaces, use of disposable loops, and avoidance of placing plate lids face-up are essential. Regular maintenance of laboratory hoods and HEPA filters further mitigates contamination risk.29
As this was a single-centre study, the generalizability of the present study findings is limited. Clinical correlation and further phenotypic and molecular characterization of the microbial isolates could have added valuable insights.
Extended incubation of subculture plates yielded an additional 3.4% clinically significant isolates. Importantly, contamination rates remained low (4.8% in blood agar vs 1.4% in MacConkey agar), supporting a safe diagnostic window for incubation of up to three completed days. Therefore, extending subculture incubation to three days represents a simple, low-cost strategy that can improve diagnostic sensitivity in laboratories relying on conventional blood culture systems. This enhanced detection has the potential to guide more accurate antimicrobial therapy and improve patient management in resource-limited settings.
ACKNOWLEDGMENTS
The authors are thankful to the Microbiology Laboratory personnel for their support in ensuring the smooth conduct of this research.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
SK and PP conceptualised the study. DK and PP collected resources. SK and MKG supervised the study and applied methodology. KC and DK performed formal analysis. PN, KC and AB performed Investigations and data curation. MKG and PP performed validation. PN wrote the manuscript. SK and DK reviewed and revised the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
ETHICS STATEMENT
The study was approved by the Institutional Ethics Committee (IEC Approval No. Dean/2024/EC/7014, dated 28.02.2024).
INFORMED CONSENT
Written informed consent was obtained from the participants before enrolling in the study.
- Chela HK, Vasudevan A, Rojas-Moreno C, Naqvi SH. Approach to Positive Blood Cultures in the Hospitalized Patient: A Review. Mo Med. 2019;116(4):313-317.
- Martinez RM, Wolk DM. Bloodstream Infections. In: Hayden RT, Wolk DM, Carroll KC, Tang YW, eds. Diagnostic Microbiology of the Immunocompromised Host. 2016:653-689.
Crossref - Rudd KE, Johnson SC, Agesa KM, et al. Global, regional, and national sepsis incidence and mortality, 1990–2017: analysis for the Global Burden of Disease Study. Lancet. 2020;395(10219):200-211.
Crossref - WHO. Sepsis. World Health Organization. March 3, 2024. Accessed December 19, 2025. https://www.who.int/news-room/fact-sheets/detail/sepsis
- Verway M, Brown KA, Marchand-Austin A, et al. Prevalence and Mortality Associated with Bloodstream Organisms: a Population-Wide Retrospective Cohort Study. J Clin Microbiol. 2022;60(4):e02429-21.
Crossref - Holmes CL, Albin OR, Mobley HLT, Bachman MA. Bloodstream infections: mechanisms of pathogenesis and opportunities for intervention. Nat Rev Microbiol. 2025;23(4):210-224.
Crossref - Vihari N, Bohra GK, Yadev RR, et al. The emergence of multidrug-resistant Gram-positive bloodstream infections in India – a single center prospective cohort study. Germs. 2023;13(3):229-237.
Crossref - Kumar A, Ellis P, Arabi Y, et al. Initiation of inappropriate antimicrobial therapy results in a fivefold reduction of survival in human septic shock. Chest. 2009;136(5):1237-1248.
Crossref - Iyer V, Castro D, Malla B, et al. Culture-independent identification of bloodstream infections from whole blood: prospective evaluation in specimens of known infection status. J Clin Microbiol. 62(3):e01498-23.
Crossref - Khan S, Das A, Mishra A, et al. Evaluation of three protocols for direct susceptibility testing for Gram-negative rods from flagged positive blood culture bottles. Microbiol Spectr. 2024;12(4):e03081-23.
Crossref - Liesman RM, Pritt BS, Maleszewski JJ, Patel R. Laboratory Diagnosis of Infective Endocarditis. J Clin Microbiol. 2017;55(9):2599-2608.
Crossref - Isaac EW, Jalo I, Difa AJ, et al. Bacterial Blood Isolates in Children: Conventional vs. Bactec Automated Blood Culture System in a Tertiary Health Centre in Gombe, North East Nigeria. Open J Med Microbiol. 2022;12(3):101-116.
Crossref - Yagupsky P. Detection of Brucellae in Blood Cultures. J Clin Microbiol. 1999;37(11):3437-3442.
Crossref - Ransom EM, Alipour Z, Wallace MA, Burnham CAD. Evaluation of Optimal Blood Culture Incubation Time To Maximize Clinically Relevant Results from a Contemporary Blood Culture Instrument and Media System. J Clin Microbiol. 2021;59(3):e02459-20.
Crossref - Kirn TJ, Weinstein MP. Update on blood cultures: how to obtain, process, report, and interpret. Clin Microbiol Infect. 2013;19(6):513-520.
Crossref - Mohus RM, Gustad LT, Furberg AS, et al. Explaining sex differences in risk of bloodstream infections using mediation analysis in the population-based HUNT study in Norway. Sci Rep. 2022;12(1):8436.
Crossref - Uslan DZ, Crane SJ, Steckelberg JM, et al. Age- and sex-associated trends in bloodstream infection: a population-based study in Olmsted County, Minnesota. Arch Intern Med. 2007;167(8):834-839.
Crossref - Anuradha S, Samaddar A, Maurya A, et al. Analysis of Blood Culture Data Influences Future Epidemiology of Bloodstream Infections: A 5-year Retrospective Study at a Tertiary Care Hospital in India. Indian J Crit Care Med. 2021;25(11):1258-1262.
Crossref - Dias SP, Brouwer MC, van de Beek D. Sex and Gender Differences in Bacterial Infections. Infect Immun. 90(10):e00283-22.
Crossref - Vasudeva N, Nirwan PS, Shrivastava P. Bloodstream infections and antimicrobial sensitivity patterns in a tertiary care hospital of India. Ther Adv Infect Dis. 2016;3(5):119-127.
Crossref - Zuza-Alves DL, Silva-Rocha WP, Chaves GM. An Update on Candida tropicalis Based on Basic and Clinical Approaches. Front Microbiol. 2017;8:1927.
Crossref - Stefanie H, Gallert C. Growth Behavior of E. coli, Enterococcus and Staphylococcus Species in the Presence and Absence of Sub-inhibitory Antibiotic Concentrations: Consequences for Interpretation of Culture-Based Data. Microb Ecol. 2016;72(4):898-908.
Crossref - Shi XX, Qiu HP, Wang J, Zhang Z, Wang YL, Sun GC. A handy method to remove bacterial contamination from fungal cultures. PLOS ONE. 2019;14(11):e0224635.
Crossref - Gumral R, Dogen A, Ilkit MM. Comparison of the contamination rates of culture media used for isolation and identification of dermatophytes. Turk J Med Sci. 2015;45(3):587-592.
- US Food and Drug Administration, Office of Regulatory Affairs, Office of Regulatory Science. Pharmaceutical Microbiology Manual. Revision 02, Document ORA.007. 2020. https://www.fda.gov/media/88801/download. Accessed October 1, 2025.
- Microbiology Society. Basic Practical Microbiology. Microbiology Society. 2019. https://microbiologysociety.org/education-outreach/resources/basic-practical-microbiology.html. Accessed October 1, 2025.
- Open Oregon Educational Resources. Microbiology Laboratory Manual: Streaking for Isolation. 2020. https://openoregon.pressbooks.pub/microbiologylaboratorymanual/. Accessed October 1, 2025.
- Kent PT, Kubica GP. Public Health Mycobacteriology: A Guide for the Level III Laboratory. Centers for Disease Control, Atlanta, GA. 1985. https://ntrl.ntis.gov/NTRL/dashboard/searchResults/titleDetail/PB86216546.xhtml. Accessed August 15, 2025.
- Nagarajan P, Mahalingam P, Aasaithambi A. Think before use: Sterility checking of culture plates. GSC Biol Pharm Sci. 2018;5(1):82-86.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.