ISSN: 0973-7510
E-ISSN: 2581-690X
The study aim to screen and characterize antibiotic producing bacteria isolated from sewage contaminated soil samples in Visakhapatnam, Andhra Pradesh, India. Sewage samples were collected and bacterial isolation was performed using serial dilution. A total of 30 bacterial isolates were obtained and they were designated as AntC1-AntC30. Each bacterial isolate was screened for its antibacterial activity following a standard agar well diffusion method against two test bacterial culture (Staphylococcus aureus and Escherichia coli). The isolate AntC3 exhibited strong antibacterial activity; hence, it was further characterized by using polyphasic taxonomic studies including 16S rRNA analysis. The potent strain showed 99% similarity to known sequences in the NCBI database and identified as Bacillus tropicus (MK396662). “Optimization of culture conditions (incubation period 18 hrs, temperature 37 °C, pH 7.0, 1% fructose, and 1% beef extract as optimal carbon and Nitrogen sources, respectively) significantly enhanced antibacterial activity”. Based on the findings from the present/current study, the strain Bacillus tropicus AntC3 isolated from the sewage samples of Visakhapatnam could be the promising source of potential bioactive metabolites.
Sewage Effluents, Bacillus tropicus AntC3, Antibacterial Activity, Optimization, Antimicrobial Activity
The misapplication of antibiotics has exhilarated the appearance of drug-resistant pathogens, and it is presently a burning issue which will gradually become a life-threatening problem for mankind. Among the notable advancements in pharmaceutical industries during the twentieth century was the development and utilization of microorganisms for antibiotic productions. Hence, it is crucial to focus on developing novel, antibiotics from microbial sources. Research on antimicrobial compounds producing from microorganisms can lead to the development of various structurally distinct antibiotics for treating microbial infections produced by drug resistant pathogens.1 Development of host resistant against the human microbial pathogens is possible by blocking the development of antibiotic resistant among the microbial pathogens with the help of active biological agents.2
Production rate of antimicrobial agents from microorganisms can be enhanced by the adjustment or optimization of the physiochemical parameters.3 The production rate of antimicrobial agents is always being found unstable, if we produced it through fermentation process, leading to unpredictable yield of the desired product. To get our desirable product at a known concentration it is possible by adjusting or optimizing certain parameters like nutrients type, pH, temperature, incubation period, etc. These parameters must show a critical impact on the production rate during production process.4 pH of the medium also very important during the production medium formulation; as it influences the expression of microbial enzymes which are involved in the production of an antibiotic.5 Research has shown that physical parameters like pH, temperature, aeration and agitation effects the microbial growth and antibiotic production rate in Xenorhabdus species.6
Progression or stagnation of the antibiotic production is possible by the optimization of the environmental factors of the culture medium or production medium. Hence optimization of suitable growth condition will promote the growth rate and production rate of an antibiotic as well. It involves an important role in the production process.7-13 Hence, the present study focused on to the isolation of bacterial species showing antimicrobial activity and furthermore optimization of the physiochemical parameters of the culture medium to enhancing the antimicrobial production.
New antibiotic producing bacteria may possibly present in sewage contaminated soil as sewage is the rich source of macromolecules and there has been an immense possibility of having novel antibiotic producers.14 However, the optimization condition of culture media of the bacterial culture which has isolated from the sewage specimens has not been documented so far. Hence, this work was initiated to give an insight into the supreme production of an antimicrobial compounds by consuming bacterial strain particularly extracted from sewage samples by using established optimization technique.
Isolation and screening of bacteria that produce antibiotics
In Visakhapatnam, Andhra Pradesh, India, the polluted soil samples were taken from the sewage effluent sites of Parawada, Madhurawada, and PM Palem. Three water samples were taken aseptically and placed in sterile glass bottles before being taken to the lab. Nutrient agar medium (NAM) was formulated and sterilized at 121 °C for 15 to 20 minutes in an autoclave in order to isolate microorganisms. Following the sterilization step, antifungal agent (Nystatin: 25 µg/ml) was added to the medium to minimize the growth of fungal contamination. Then the prepared medium was poured into Petri plates and medium was allowed for solidification. Serial dilution technique was employed for isolating the bacteria; 0.1 ml suspension from each dilution (10-6-10-7) was spread over NAM plates. Triplicates of each sample were made and they were incubated for 48 hours at 35 ± 2 °C. Following incubation, appeared bacterial colonies were selected, sub-cultured and preserved on NAM slants at 4 °C for further studies.9 Identified isolates were screened for the antimicrobial activity through agar well diffusion technique against Staphylococcus aureus and Escherichia coli.
Identification of potent strain by Polyphasic taxonomy
The cultural characteristics of the potent strain were examined by inoculating it on to nutrient agar plates aseptically and incubated at 35 ± 2 °C for 2-3 days. Colony characteristics and pigment production were morphologically observed. Slides were prepared for Gram’s staining and observed under the microscope at magnification of ×100.10 Biochemical characterization of AntC3 bacterial strain was performed using Indole, Citrate utilization, methyl red and Voges-Proskauer tests.11
16S rRNA analysis for strain identification
The genomic DNA of bacterial strain was isolated using the InstaGeneTM matrix kit (Bio-Rad, USA). Few bacterial colonies were collected, suspended in sterile water (1 ml), and centrifuged (10,000-12,000 rpm for 1 minute) to remove the supernatant. The pellet was then treated with InstaGene matrix (200 µl), incubated at 56 °C for 15 minutes and then in a boiling water bath at 100 °C for 8 minutes, vortexing with high speed (10 seconds) in between steps. The sample was then centrifuged (at 10,000-12,000 rpm about 2 minutes), and the DNA rich supernatant was collected after centrifugation, with a 20 µL sample used for PCR.13 The 16S rRNA gene amplification was performed for the genomic DNA template using thermal cycler and universal primers, 27F-AGAGTTTGATCMTGGCTCAG, 1492R-TACGGYTACCTTGTTACGACTT. The PCR reaction, a 20 µL solution containing 1 µL of template DNA, was set for an initial denaturation at 94 °C for 2 minutes The 35 amplification cycles each consisted of denaturation (at 94 °C for 45 seconds), annealing (at 55 °C for 60 seconds), and extension (at 72 °C for 60 seconds); followed by a final extension at 72 °C for 10 minutes This resulted in the amplification of 1400 bp DNA fragments. Both Negative and positive control (E. coli genomic DNA) were included in the PCR reaction.14
Bioinformatics tools
NCBI Nucleotide BLAST program was used to identify the sequences by uploading FASTA a similarity searching tool. Forward and reverse sequences were aligned by using the SequentiX software DNA Dragon and prepare FASTA. The multiple sequence alignments were performed using MUSCLE 3.7 program, followed by Phylogenetic tree construction using the MEGAX 5.2 software.
Growth pattern and impact of incubation period on antimicrobial production
To investigate the growth pattern and antimicrobial activity of Bacillus tropicus AntC3, a study was conducted over a 22 hour period, with samples taken every 2 hours. A seed culture was first prepared by inoculating the strain into 100 ml of Nutrient broth and incubating it at 30 ± 2 °C and 120 rpm for 24 hours. A 1 ml aliquot of this culture was then used to inoculate a fresh 100 ml batch of Nutrient broth, which was incubated under the same conditions. At each 2 hour interval, the biomass was separated from the broth. The remaining culture filtrate was extracted with an equal volume of ethyl acetate (1:1). The solvent was removed from the extract using a rotary evaporator under vacuum, and the resulting residue was tested for antibacterial activity by agar well diffusion. The effectiveness was measured by the diameter of the inhibition zones against two indicator organisms: Staphylococcus aureus and Escherichia coli.
Impact of carbon and nitrogen sources on antimicrobial production
The impact of different carbon sources on antimicrobial compounds production by Bacillus tropicus AntC3 was studied by adding various carbon sources viz., galactose, mannitol, fructose, glucose, maltose and starch to the Nutrient broth at 1% concentration, replacing the existing carbon source.15 The optimized carbon source was used further.
Similarly, the impact of different nitrogen sources were studied by supplementing the Nutrient broth with 1% of Ammonium nitrate, ammonium sulphate, yeast extract, beef extract and peptone by replacing the standard nitrogen sources.15 The optimized nitrogen source that resulted in enhanced antimicrobial compound production was identified and used for further studies.
Impact of pH and temperature on antimicrobial production
The adjustment of the pH of production medium between 5.0 to 9.0 (using 1 M HCL and 1 N NaOH) determined the specific pH at which antimicrobial compounds production is maximized, as evidenced by the largest zone of inhibition and used as reference point for further study.
The optimal temperature was evaluated by incubating the inoculated production medium at three different temperatures (28 °C, 37 °C and 42 °C). The culture filtrates were then extracted using a solvent extraction method by observing the zone of inhibition and identified the optimal temperature.
Isolation and screening of bacteria that produce antibiotics
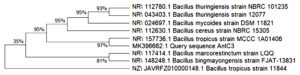
A total of thirty bacterial isolates, labeled as AntC1-AntC30 were isolated from sewage effluent contaminated soil samples collected in Visakhapatnam. In screening, three isolates such as AntC1, AntC3 and AntC8 were found to be potent and exhibited antibacterial activity (Table). Among them AntC3 exhibited highest antibacterial potential against Staphylococcus aureus and Escherichia coli. The potential bacterial isolate (AntC3) was identified using the methods outlined in Bergey’s Manual of Systematic Bacteriology. In micro-morphological analysis revealed that AntC3 isolate was a Gram-positive, rod shaped Bacteria with varied size. Biochemical tests gave negative results for Indole, Methyl-red and positive results for both citrate utilization, Voges-Proskauer tests. Molecular characterization using 16S rRNA analysis identified the AntC3 isolate as Bacillus tropicus. The identified partial sequence was deposited in the NCBI. Genbank database, obtained the accession number MK396662. The Phylogenetic tree (Figure 1) was constructed with the Neighbor-joining (NJ) method in MEGA-X, Version 5.0 software.
Table:
Antibacterial activity of the bacterial isolates
| No. | Bacterial isolates | Zone of inhibition (mm) | |
|---|---|---|---|
| S. aureus | E. coli | ||
| 1 | AntC1 | 10 ± 1 | 02 ± 1 |
| 2 | AntC3 | 17 ± 1 | 20 ± 1 |
| 3 | AntC8 | 08 ± 1 | 09 ± 1 |
Figure 1. Phylogenetic analysis of AntC3 and closely related Bacillus species based on 16S rRNA gene Sequence by Neighbor joining method
Growth pattern and impact of incubation period on antimicrobial production
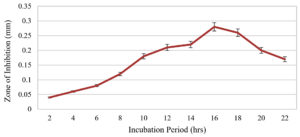
The strain Bacillus tropicus AntC3 growth pattern and antimicrobial property were studied at regular intervals up to 22 hours in batch culture. The bacterial strain entered into log phase after 4 hours of incubation. It entered the stationary phase after 16 hours of incubation which extended up to 18 hours (Figure 2). The secondary metabolites produced from 18 hours old culture exhibited high antibacterial effect against the Escherichia coli, as evidenced by 14 ± 2 mm inhibition zone and Staphylococcus aureus with a corresponding inhibition zone of 16 ± 2 mm mostly the different Bacillus strains require varying incubation period for the production of antimicrobial metabolites. Comparatively, the optimal incubation periods for bioactive metabolite production varied among Bacillus strains. For instance, Bacillus safensis MK 12.116 and Bacillus pumilus17 showed high antimicrobial activity after 48 hours, while Bacillus amyloliquefaciens BS418 exhibited optimal activity after 24 hours. Conversely, Bacillus subtilis RLID 12.119 demonstrated maximum antimicrobial production after just 5 hours.
Impact of carbon and nitrogen sources on antimicrobial production
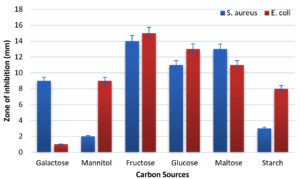
The impact of different carbon source on antimicrobial compound production of Bacillus tropicus AntC3 strain is shown in Figure 3. In the Nutrient broth medium supplemented with 1% of fructose yielded the highest secondary metabolite production, followed by 1% of Glucose and Maltose (Figure 3). The zone of inhibition for each carbon source was evaluated individually. Fructose exhibited the largest inhibition zone, with 14 ± 1 mm against Staphylococcus aureus and 15 ± 1 mm against Escherichia coli. Glucose and maltose also showed significant zones of inhibition, with 11 ± 1 mm and 13 ± 1 mm against S. aureus, and 13 ± 1 mm and 11 ± 1 mm against E. coli, respectively. While Galactose and Mannitol showed moderate growth of inhibition of 9 ± 1 mm of S. aureus, and E. coli respectively and showed low zone of inhibition for Galactose (1 ± 0.1 mm) and for Mannitol (2 ± 0.1 mm) effect on E. coli and S. aureus respectively. Similarly moderate growth inhibition measuring 8 ± 2 mm against E. coli and low inhibition zone 3 ± 1 mm against S. aureus was observed in presence of starch. It was evidently reported that fructose is the suitable carbon source to enhance the antimicrobial activity production in Bacillus circulans NB-7.15 In contrast, Semolina flour was proved as an excellent carbon source for enhancing antibiotic production by the strain Bacillus amyloliquefaciens BAM.20
Figure 3. Impact of carbon sources on antimicrobial production by Bacillus tropicus AntC3
(The Values are the mean of three replicates standard Deviation (± SD) and were found to be significant at 5% level)
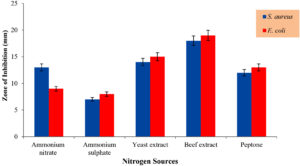
To identify the most effective nitrogen source for promoting secondary metabolite production, various nitrogen sources were evaluated. Among the tested sources, beef extract (1%) emerged as the optimal choice, yielding the largest zones of inhibition: 18 ± 1 mm against Staphylococcus aureus and 19 ± 1 mm against Escherichia coli (Figure 4). Yeast extract also showed significant zones of inhibition, with 14 ± 2 mm for S. aureus and 15 ± 1 mm for E. coli. Peptone showed moderate zones of inhibition, with 12 ± 1 mm for S. aureus and 13 ± 1 mm for E. coli. In contrast, ammonium nitrate and ammonium sulphate showed relatively lower zones of inhibition, with 13 ± 1 mm and 9 ± 2 mm of S. aureus and E. coli, and with 7 ± 1 mm and 8 ± 1 mm of S. aureus and E. coli, respectively. Interestingly in a previous study, beef-extract shown to be a more appropriate Nitrogen source for enhancing antimicrobial production in Bacillus amyloliquefaciens BAM strain than other alternate sources.20
Figure 4. Impact of Nitrogen sources on antimicrobial production by Bacillus tropicus AntC3
(The Values are the mean of three replicates standard Deviation (±SD) and found to be significant at 5% level)
Impact of pH and temperature on antimicrobial production
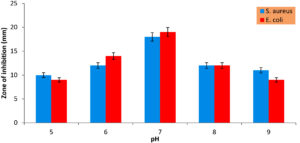
The impact of pH and temperature on secondary metabolite production of the strain is illustrated in Figures 5 and 6. Optimal bioactive metabolite production was achieved at pH 7.0, resulting in the largest zones of inhibition: 18 ± 1 mm against Staphylococcus aureus and 19 ± 1 mm against Escherichia coli. In contrast, acidic conditions (pH 5.0) yielded relatively lower zones of inhibition, measuring 10 ± 1 mm against S. aureus and 9 ± 1 mm against E. coli. At pH 6.0, the zones of inhibition were 12 ± 1 mm against S. aureus and 14 ± 2 mm against E. coli. Similarly, at pH 8.0, the zones of inhibition remained consistent, measuring 12 ± 1 mm against both S. aureus and E. coli. However, at pH 9.0, the zones of inhibition decreased to 11 ± 1 mm against S. aureus and 9 ± 1 mm against E. coli. Interestingly, similar trends have been observed in other Bacillus species, such as Bacillus cereus, Bacillus endophyticus, and Bacillus subtilis, which also exhibit enhanced antibiotic production at pH 7.0.21
Figure 5. Impact of pH on antimicrobial production
(The Values are the mean of three replicates standard Deviation (±SD) and found to be significant at 5% level)
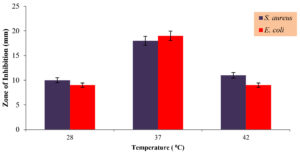
Figure 6. Impact of temperature on antimicrobial production
(Values are the means of three replicates standard Deviation (±SD) and found to be significant at 5% level)
The strain showed an optimal antimicrobial production at 37 °C which was determined with an inhibition zone of 18 ± 1 mm and 19 ± 1 mm against Staphylococcus aureus and Escherichia coli respectively, hence the strain was appeared to be mesophilic. Deviation from this optimal temperature resulted in decrease the production of antimicrobial compound. This finding aligns with earlier reports on various Bacillus species.22 including Bacillus subtilis RLID 12.1,19 Bacillus endophyticus RN-7.23 which also exhibit maximal bioactive metabolites production at 37 °C.22
A potent antimicrobial agent, identified as Bacillus tropicus AntC3 (MK396662), was successfully isolated from sewage effluents in Visakhapatnam, Andhra Pradesh, India. Notably, this strain exhibited strong antimicrobial property against with both Staphylococcus aureus and Escherichia coli. Optimization of growth conditions (pH of 7.0, 37 °C temperatures, 1% fructose, and 1% beef extract) maximized the bioactive metabolite production. Future studies focusing on further characterization and purification of these metabolites will provide valuable insights into their potential pharmaceutical applications.
ACKNOWLEDGMENTS
The authors sincerely acknowledged the Department of Botany and Microbiology, Acharya Nagarjuna University, and MIC Gene laboratory, Visakhapatnam, for providing the laboratory facilities.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
JM conceptualized the study. VVSRKK performed literature review, experiments and wrote the manuscript. JM reviewed the manuscript. Both authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies on human participants or animals performed by any of the authors.
- Durand GA, Raoult D, Dubourg G. Antibiotic discovery: history, methods and perspectives. Int J Antimicrob Agents. 2019;53(4):371-382.
Crossref - Wright GD. Antibiotic adjuvants: rescuing antibiotics from resistance. Trends Microbiol. 2016;24(11):862-871.
Crossref - Djinni I, Defant A, Kecha M, Mancini I. Metabolite profile of marine derived endophytic Streptomyces sundarbansensis WR 1 L 1 S 8 by liquid chromatography-mass spectrometry and evaluation of culture conditions on antibacterial activity and mycelial growth. J Appl Microbiol. 2014;116(1):39-50.
Crossref - Kiranmayi MU, Sudhakar P, Sreenivasulu K, Vijayalakshmi M. Optimization of culturing conditions for improved production of bioactive metabolites by Pseudonocardia sp. VUK-10. Mycobiology. 2011;39(3):174-181.
Crossref - Gao X, He Q, Jiang Y, Huang L. Optimization of nutrient and fermentation parameters for antifungal activity by Streptomyces lavendulae Xjy and its biocontrol efficacies against Fulvia fulva and Botryosphaeria dothidea. J Phytopathol. 2016;164(3):155-165.
Crossref - Webster JM, Chen G, Hu K, Li J. Bacterial metabolites. Entomopathog Nematol. 2002;99-114.
Crossref - Gopi R, Ramakrishna R, Rajagopal A. Optimization of culture conditions of Streptomyces rochei (MTCC 10109) for the production of antimicrobial metabolites. Egypt J Biol. 2011;13(1):21-29.
Crossref - Aman GZ. Studies on Some Actinomycete Isolate from Certain Desert Soil Samples. Ph.D. Thesis. Faculty of Science. Egypt: Al-Azhar University. 2001.
- Jonsbu E, McIntyre M, Nielsen J. The influence of carbon sources and morphology on nystatin production by Streptomyces noursei. J Biotechnol. 2002;95(2):133-144.
Crossref - Padma PN, Rao AB, Yadav JS, Reddy G. Optimization of fermentation conditions for production of glycopeptide antibiotic vancomycin by Amycolatopsis orientalis. Appl Biochem Biotechnol. 2002;102- 103(1-6):395-405.
Crossref - El-Naggar MY, Hassan MA, Said WY, El-Aassar SA. Effect of support materials on antibiotic MSW2000 production by immobilized Streptomyces violatus. J Gen Appl Microbiol. 2003;49(4):235-243.
Crossref - Sharon SF, Daniel RR, Shenbagarathai R. Optimization of antibiotic production by marine actinomycetes Streptomyces sp. Int J Pharm Pharm Sci. 2014;6(8):506-510.
- Kiviharju K, Leisola M, Eerikainen T. Optimization of Streptomyces peucetius var. caesius N47 cultivation and epsilon-rhodomycinone production using experimental designs and response surface methods. J Ind Microbiol Biotechnol 2004;31(10):475-481.
Crossref - Russell AD, Furr JR. The antibacterial activity of a new chloroxylenol preparation containing ethylenediamine tetra acetic acid. J Appl Bacteriol. 1977;43(2): 253-260.
Crossref - El-Banna NM. Effect of carbon source on the antimicrobial activity of the air flora. World J Microbiol Biotechnol. 2005;21(8):1451-1454.
Crossref - Iqbal S, Begum F, Qasim M. Screening, Characterization and Optimization of Bioactive Peptides with Antibacterial Activities Against Multi-Drug Resistant Pathogens, Produced by Bacillus safensis Strain MK-12.1. Int J Pept Res Ther. 2022;28(6).
Crossref - Amin A, Khan MA, Ehsanullah M, Haroon U, Azam SMF, Hameed A. Production of peptide antibiotics by Bacillus sp: GU 057 indigenously isolated from saline soil. Braz J Microbiol. 2012;43(4):1340-1346.
Crossref - Palacios-Rodriguez AP, Espinoza-Culupu A, Duran Y, Sanchez-Rojas T. Antimicrobial Activity of Bacillus amyloliquefaciens BS4 against Gram-Negative Pathogenic Bacteria. Antibiotics. 2024;28:13(4):304.
Crossref - Ramachandran R, Chalasani AG, Lal R, Roy U. A Broad-Spectrum Antimicrobial Activity of Bacillus subtilis RLID 12.1. Sci World J. 2014:968487.
Crossref - Ahsan T, Zang C, Yu S, et al. Screening, and optimization of fermentation medium to produce secondary metabolites from Bacillus amyloliquefaciens, for the biocontrol of early leaf spot disease, and growth promoting effects on Peanut (Arachis hypogaea L.). J Fungi. 2022;8(11):1223.
Crossref - Nagaraju K, Ram MR. Screening and Identification of L-glutaminase Producing Bacteria From Forest Soils. Int J Pharm Res. 2018;10(2):66-69.
- Neelam B, Gupta RK. Effect of pH and Temperature on Secondary Metabolite Isolated from Soil Bacteria. Int J Life Sci Sci Res. 2018;4(2):1729-1735.
Crossref - Nagaraju K, Ram MR. Studies on Optimization of L-Glutaminase from Bacillus Endophyticus RN-7. Int J Pharm Biol Sci. 2018;8(3):11746-1153.
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.