ISSN: 0973-7510
E-ISSN: 2581-690X
Helicobacter pylori, a Gram-negative gastric pathogen and a class I carcinogen harbors key virulence genes such as vacA, cagA, ureC, homB and oipA that contributes to gastrointestinal diseases. The study assessed the prevalence of H. pylori infection and their correlation with virulence genotypes with clinical outcomes in South Indian Tamils. Gastric antral biopsies from 500 patients were assessed for H. pylori prevalence by RUT followed by genomic DNA extraction and PCR. Molecular confirmation employed 16S rRNA, a preliminary marker, with the identification of virulent genes cagA, vacAs1, vacA m1/m2, oipA, ureC, homB are validated by sequencing. H. pylori were confirmed in 491 patients where vacA s1 (71.89%) was correlated with gastritis (OR-1.68;95%; CI: 1.11-2.53; p = 0.013) and pangastritis (OR-4.79;95%; CI: 2.02-11.34; p = 0.0004) while vacA s1/m1 and vacA s1/m2 alleles were found to be 51.53% and 14.26%, respectively. Gastritis was associated with s1/m1 (OR-2.34; 95%CI: 1.29-4.23; p = 0.004), s1/m2 (OR-2.03; 95%CI: 1.04-3.95; p = 0.03) and pangastritis (OR-2.61;95%CI: 0.97-7.02; p = 0.05) was associated with s1/m1. The cagA gene (31.36%) was correlated to gastric cancer (OR-3.45; 95%CI: 1.20-9.89; p = 0.02) and oipA gene (64.97%) was associated with PUD (OR = 7.87; 95% CI: 1.02–60.40; p = 0.04), homB (24.03%) with gastritis (OR-1.62; 95%CI: 1.01-2.59; p = 0.04) and ureC (59.67%) with no disease association. Therefore, our study provides genotypic prevalence of H. pylori infection in the South Indian Tamil population, with vacA s1 being the predominant genotype reported and significantly associated with gastritis and pangastritis. Hence, vacA s1 can serve as a potent virulent marker for gastrointestinal disease manifestations.
Gastric Cancer, Gastritis, Helicobacter pylori, PUD, South Indian Population, vacA
Helicobacter pylori is a Gram-negative bacterium that typically colonizes in the antral region of the human stomach. The pathogen was considered to be a class I carcinogen, as classified by the WHO, and was associated with various gastric complications such as gastritis, peptic ulcer disease (PUD), dyspepsia, mucosa-associated lymphoid tissue (MALT) lymphoma and gastric cancer (GC).1 The frequency of H. pylori infection in Asia was reported high, reaching almost 80% of the population with the prevalence rate surpassing 50% by the age of 10, thereby rising to 90% in adulthood, while in developed countries like Japan, Western Europe and Oceania, it was estimated to be between 30% and 50%. Moreover, a recent meta-analysis report among 15 different states of Indian population have shown a marginal regional variation in the prevalence of infection among the states, where Rajasthan reports highest prevalence of 70% and the lowest prevalence was reported in Gujarat (9%). Also, the report has analyzed the pooled prevalence of H. pylori among people with GI disorders was 54% in comparison with 61% of no clinically diagnosed GI disorders.2-4 Studies have reported that the recommended initial treatment for H. pylori infection includes a seven-day vonoprazan-based triple therapy and a non-bismuth quadruple therapy for about ten to fourteen days. These regimens have achieved eradication rates of about 90%, even in regions with high prevalence of antimicrobial-resistant strains. Nonetheless, these treatments still have certain limitations, such as the potential to increase antimicrobial resistance and causing imbalances in gut microbiota.5 The pathogenesis and adverse effects of the disease are influenced by complex interactions between bacterial virulence, host responses, and environmental factors. Over time, these virulence factors contribute to the persistence and colonization of H. pylori. The recognized virulence factors of H. pylori are vacuolating cytotoxin A (vacA), induced by contact with the epithelium (iceA), outer membrane proteins such as helicobacter outer membrane proteins (hom), outer inflammatory protein A (oipA), urease (ure), cagPAI (cag pathogenicity island) and blood group antigen-binding adhesion (babA) which have the potential to interfere with gastric epithelial cell signaling pathways thereby inducing gastric diseases.6-8
cag PAI of H. pylori, comprising 40 kb of DNA, housing for 27-31 genes, a significant virulence factor which codes for cagA has been identified in adverse cases of gastrointestinal ailments, such as acute gastritis, PUD and GC. The cagA gene represents part of the type IV secretion system (T4SS), which acts as translocation complex to deliver the cagA protein into the cytoplasm of the infected gastric cells, triggering cytoskeletal changes, increased cell proliferation, and release of interleukin-8 (IL-8).9 vacA, a 140 kDa pore-generating toxin, encoded by a 3.9 kb gene and induces the formation cytoplasmic vacuoles, mitochondrial fragmentation, permeabilization of cell membranes and kinase enzyme activation. The vacA gene has two allelic forms namely, signal peptide (s1, s2), and middle region (m1, m2). Moreover, the combination of the allelic forms also known to cause higher levels of toxins as produced by s1/m1 type strains in comparison to s2/m2 are associated with gastric cancer and premalignant conditions. It has been reported that the vacA s1m1 genotype was reported to be highly dangerous because of the high level of toxin production, secretion of inflammatory chemicals, and an increased risk of GC.10 The oipA gene plays a vital role in triggering IL-8 secretion and enhancing bacterial invasion in the stomach.11 The regulation of oipA gene was mediated via slipped-strand mispairing, which is contingent on the number of CT repeats. The expression of this toxin leads to apoptosis, cellular stress, and alteration in cytoskeletal of gastric epithelial cells, causing significant gastrointestinal disorder.12 The ureC gene encodes an open reading frame that originated from the urease structural gene (ureAB) of H. pylori, a phosphoglucosamine mutase derivative.13 An investigation indicates that the gene has been strongly associated with gastric cancer.14 The hom family represents a small group of paralogous proteins characterized by signal sequences in the C-terminal and interchanging hydrophobic motifs. The homA and homB genes are 90% identical with a variation in the central domain. Recent studies reported that homB has a close association with IL-8 secretion and a significant association with gastritis, PUD and gastric cancer.15
In 2022, India was ranked third globally for stomach cancer incidence and second for gastric cancer mortality.16 The recent metanalysis study on H. pylori infection highlighted a significant regional disparities, among various parts of India with prevalence ranging from 9%-70%.4 Based on the higher prevalence of H. pylori infection in India and its well-established role in gastrointestinal diseases, our study aims to investigate the genotypic profile of H. pylori virulent strains isolated from gastric biopsy sample of South Indian Tamil population. By determining the prevalence of key virulence genes and analyzing their association with specific gastrointestinal conditions, our study will provide the insights into strain-specific risks. Also, the findings of our study may enhance the understanding of disease progression to aid in the identification of molecular markers for detection and targeted therapeutic interventions.
Study subjects and sample collection
The Institutional Ethical Committee has granted approval for the proposed study (EC Reg No. 2394/IEC/2021) and informed consent was acquired from the H. pylori infected patients prior to their recruitment. Also, the committee has granted approval for study objective which demands patients tested positive for H. pylori during the endoscopic procedure and restricted the inclusion of a true H. pylori negative control group. Blood and gastric biopsy tissues were procured from 500 patients who underwent upper gastrointestinal endoscopy at SRM Medical College and Research Centre, Tamil Nadu, India. H. pylori infection was confirmed through the rapid urease test (RUT) and ELISA to determine IgG levels against the bacteria (HPIgG) (Data not shown). A cutoff value of ≥30 U/mL was considered positive for active H. pylori infection.17 Two biopsy samples were taken from the patient during the endoscopic procedure: one for histopathological study and the other for further molecular studies. The biopsy tissue samples were transported to the laboratory under ice-cold conditions in a transport medium containing 0.6 ml of Brucella broth and 15% glycerol and stored at -80 °C until further analysis.18
Investigation of biopsy samples
Rapid Urease Test
The tissue samples collected from each subject were placed on an RUT strip purchased from Gastrohub in Kolkata, India. Two drops of sterile water were added, and was observed for the colour change from yellow to pink or red within 2 h was considered positive. The test has detected the presence of H. pylori infection through its urease activity.19
H. pylori confirmation by histopathology
Histopathologic confirmation was performed by fixing the biopsy in 10% formalin, embedded in paraffin, sectioned and stained using giemsa staining solution to detect spiral or rod-shaped bacteria on the gastric epithelium.20
DNA extraction and genotyping of H. pylori virulence genes
Genomic DNA was extracted using the QIAamp® DNA Mini kit, from the stomach tissues according to the manufacturer’s protocol and stored in -20 °C until use. The genotyping analysis was performed with 16S rRNA and gene-specific primers with the PCR cycling conditions mentioned in Table 1.21-26 PCR was performed in a 20 µl reaction mixture containing 10 µl Taq DNA Polymerase 2x Master Mix RED, 2 µl (10-50 ng) of genomic DNA, 1 µl of 10 pm forward and reverse primers for vacA s1, m1/m2, oipA, ureC, homB and 2 µl of 2.5 pm primers of cagA and the rest nuclease-free water. PCR products were visualized on 2% agarose gel stained with ethidium bromide. The clinical isolates were subjected to nucleotide sequencing analysis to confirm the genotyping assay.
Table (1):
Primer sequences and PCR conditions of H. pylori 16S rRNA, virulent genes and their PCR product size
Genes |
Primer sequence (5ʹ→3ʹ) |
PCR Condition |
Size of PCR Product (bp) |
Ref. |
|---|---|---|---|---|
16S rRNA |
5’-GCGCAATCAGCGTCAGGTAATG-3’ 5′-GCTAAGAGAGCAGCCTATGTCC-3′ |
Denaturation for 30 Secs @ 94 °C; Annealing for 30 Secs @ 53 °C and extension for 45 Secs @ 72 °C (40 cycles) |
522 bp |
22 |
vacA s1 |
5’-ATGGAAATACAACAAACACAC-3’ 5’-CTGCTTGAATGCGCCAAAC-3’ |
Denaturation for 30 Secs @ 94 °C; Annealing for 30 Secs @ 55 °C and extension for 40 Secs @ 72 °C (35 cycles) |
259 bp |
23 |
CagA |
5’-GTTGATAACGCTGTCGCTTCA-3’ 5’-GGGTTGTATGATATTTTCCATAA-3’ |
Denaturation for 30 Secs @ 94 °C; Annealing for 30 Secs @ 55 °C and extension for 40 Secs @ 72 °C (35 cycles) |
351 bp |
24 |
vacA m1/m2 |
5’-CAATCTGTCCAATCAAGCGAG-3’ 5’-GCGTCAAAATAATTCCAAGG-3’ |
Denaturation for 30 Secs @ 94 °C; Annealing for 30 Secs @ 56 °C and extension for 30 Secs @ 72 °C (35 cycles) |
587 bp/642 bp |
23 |
ureC |
5’-CCCTCACGCCATCAGTCCCAAAAA-3’ 5’-AAGAAGTCAAAAACGCCCCAAAAC-3’ |
Denaturation for 30 Secs @ 94 °C; Annealing for 30 Secs @ 63 °C and extension for 40 Secs @ 72 °C (35 cycles) |
413 bp |
25 |
homB |
5’-AGAGGGTGTTTGAAACGCTCAATA-3’ 5’-GGTGAATTCTTCTGCGGTTTG-3’ |
Denaturation for 30 Secs @ 94 °C; Annealing for 30 Secs @ 63 °C and extension for 30 Secs @ 72 °C (35 cycles) |
161 bp |
21 |
oipA |
5’-CAAGCGCTTAACAGATAGGC-3’ 5’-AAGGCGTTTTCTGCTGAAGC-3’ |
Denaturation for 30 Secs @ 94 °C; Annealing for 30 Secs @ 63 °C and extension for 40 Secs @ 72 °C (35 cycles) |
428 bp |
26 |
Statistical Analysis
All the statistical analysis were performed using IBM SPSS Software (version 23). The Chi-test (χ2) was used to analyze significant differences in the demographic characteristics (sex, age, smoking/alcohol and tobacco consumption) in our study population. Odds Ratios (ORs) with 95% confidence intervals (CIs) were calculated to evaluate the link between gastrointestinal diseases and the presence of virulence genes with statistical significance at p-values ≤0.05.
Confirmation of H. pylori from biopsy samples
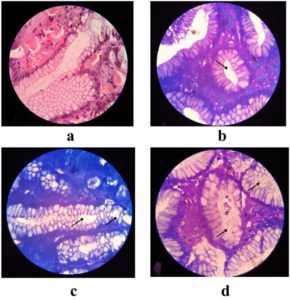
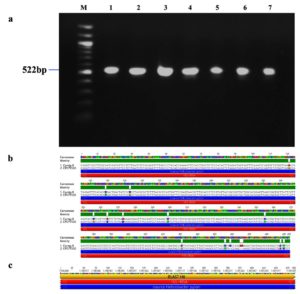
The study examined 500 patients who underwent H. pylori detection via RUT, correlated with various gastrointestinal disorders (Table 2). Giemsa staining was performed on the gastric biopsies, confirming the presence of rod-shaped bacilli within the gastric epithelial cells, as illustrated in Figures 1a-d, revealing H. pylori infection. Further confirmation involved PCR analysis for the identification of the 16S rRNA gene specific to H. pylori which showed amplification at 522 bp in 491 samples (Figure 2a). The sequence of the 16S rRNA gene was subsequently analyzed using Geneious Software showing 97.33% similarity to the reference strain (CP079530 NCBI), with minor variation as shown in Figure 2b and Figure 2c. The demographic analysis revealed that 279 participants were male with a mean age of 47.16 ± 15.74 year, while 221 were female with a mean age of 47.87 ± 14.35 years. Also, the social behaviour of both the groups was analyzed and we identified a significant association between smoking/alcohol, tobacco consumption and H. pylori, as shown in Table 3.
Table (2):
Particulars of endoscopic findings of H. pylori-infected cases based on admission diagnosis and their percentage prevalence of virulence genes associated
Endoscopic Findings (n = 500) |
PCR Positive for 16S rRNA |
cagA Positive (%) |
vacA s1 Positive (%) |
ureC Positive (%) |
oipA Positive (%) |
vacA m1 Positive (%) |
vacA m2 Positive (%) |
homB Positive (%) |
vacA s1/m1 Positive (%) |
vacA s1/m2 Positive (%) |
|---|---|---|---|---|---|---|---|---|---|---|
Gastritis (339) |
333 |
96 (19.5) |
251 (51.1) |
199 (67.9) |
217 (68.2) |
175 (69.2) |
73 (70.2) |
89 (75.5) |
124 (70.1) |
52 (74.3) |
Duodenal Ulcer (36) |
36 |
12 (2.4) |
21 (4.2) |
24 (8.2) |
30 (9.4) |
21 (8.3) |
9 (8.7) |
2 (1.7) |
11 (6.2) |
4 (5.7) |
Gastric cancer (9) |
15 |
9 (1.8) |
4 (0.8) |
11 (3.8) |
4 (1.3) |
3 (1.2) |
2 (1.9) |
3(2.5) |
3 (1.7) |
2 (2.9) |
Gastric ulcer (15) |
9 |
5 (1.01) |
2 (0.4) |
6 (2.05) |
9 (2.83) |
3 (1.19) |
4 (3.85) |
0 |
1 (0.56) |
0 |
Pangastritis (82) |
79 |
23 (4.6) |
66 (13.4) |
42 (14.3) |
42(13.2) |
41 (16.2) |
14 (13.5) |
23 (19.5) |
33 (18.6) |
12 (17.1) |
PUD (4) |
15 |
7 (1.4) |
6 (1.2) |
9 (3.07) |
14 (4.4) |
9 (3.5) |
2 (1.9) |
1 (0.8) |
4 (2.2) |
0 |
Nodular Gastritis (15) |
4 |
2 (0.4) |
3 (0.6) |
2 (0.6) |
3 (0.9) |
1 (0.4) |
0 |
0 |
1 (0.6) |
0 |
Total (%) |
491 |
154 (31.36) |
353 (71.89) |
293 (59.6) |
319 (64.97) |
253 (51.52) |
104 (21.18) |
118 (24.03) |
177 (36.05) |
70 (14.26) |
Endoscopic findings (n = 500), PCR confirmation by 16s rRNA specific for H. pylori (n = 491) and virulence genes prevalence was calculated from PCR-positive isolates
Table (3):
Demographic characteristics of patients upon confirmation by 16S rRNA specific for H. pylori
Variable |
Subjects n (%) |
H. pylori Positive n (%) |
H. pylori Negative n (%) |
p-value |
|---|---|---|---|---|
All |
500 (100) |
491 (98.2) |
9 (1.8) |
|
Sex |
0.988 |
|||
Male |
279 (55.8) |
274 (55.8) |
5 (55.6) |
|
Female |
221 (44.2) |
217 (44.2) |
4 (44.4) |
|
Age |
0.797 |
|||
less than 35 |
133 (26.6) |
131 (26.7) |
2 (22.2) |
|
35-45 |
111 (24.4) |
108 (22) |
3 (33.3) |
|
46-60 |
152 (30.4) |
149 (30.3) |
3 (33.3) |
|
>60 |
104 (20.8) |
103 (21) |
1 (11.1) |
|
Smoking/Alcohol |
0.053* |
|||
Yes |
96 (19.2) |
94 (18.8) |
2 (0.4) |
|
No |
404 (80.8) |
397 (79.4) |
7 (1.4) |
|
Tobacco |
0.018* |
|||
Yes |
49 (9.8) |
48 (9.6) |
1 (0.2) |
|
No |
451 (90.2) |
443 (88.6) |
8 (1.6) |
n (%), represents the number (percentage) of patients with the demographic conditions *Chi-square test was performed to compare demographic variables and patients with a positive result for H. pylori. Highlighted values denote a statistically significant result at the p < 0.05
Figure 1. Histopathological images from clinical isolates representing the level of infection by H. pylori, with black colour arrow representing the colonization of H. pylori (a) Normal Gastric mucosa (b) Mild infection by H. pylori (c) Moderate infection by H. pylori (d) Severe infection by H. pylori
Figure 2. Identification and sequence analysis of 16S rRNA gene from H. pylori isolated biopsies. (a) Agarose gel electrophoresis of 1% agarose to identify 16S rRNA gene amplified at 522 bp from H. pylori isolated biopsies where M represents 100 bp ladder and lanes represent the positive samples. (b) Pairwise identity and nucleotide sequence for 16S rRNA compared with the standard strain: CP079530 (NCBI) with 97.33% identity (c) Blast Hit of 16S rRNA amplified region from 1,186,868 →1,187,357, relative to the original gene of the reference strain
Genotypic status of virulence genes of H. pylori in different clinical manifestations
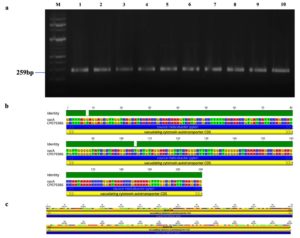
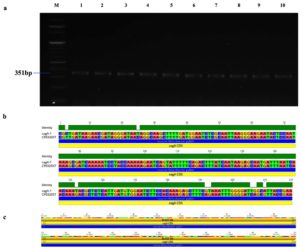
Genotypic study of 491 H. pylori isolates revealed distinct prevalence patterns for key virulence genes as described in Table 2. The vacA s1 gene exhibited the highest prevalence at 71.89% (353 of 491; p = 0.0003) of samples (Figure 3a), showing 99.2% similarity and isolates varying from the reference strain CP079377 (Figure 3b and Figure 3c). The cagA gene was found in 31.36% (154/491; p = 0.199) of the patients, as shown in Figure 4a, and exhibited 96.33% identity (Figure 4b), though variations existed among isolates when compared to the CP032037 standard (Figure 4c). Amplification of the vacA m1/m2 gene was shown in Figure 5a., accounted for approximately 51.52% (253/491; p = 0.0003) of vacA m1 and 21.18% (104/491; p = 0.044) of vacA m2 gene strains and showed a 91.2% similarity and variability against their reference (CP078468) (Figure 5b and Figure 5c). While Figure 6a shows the amplification of the ureC gene, with 59.6% (293/491; p = 0.727) of the samples being positive, which showed 97.5% similarity and variation in comparison to strain CP079049 (Figures 6b-c) The oipA gene of H. pylori was detected in 64.97% (319/491; p = 0.0003) of the patient cohort (Figure 7a), showing 90.8% similarity variations compared to its reference (CP122953) as depicted in Figure 7b and Figure 7c. The homB was identified in 24.03% (118/491; p = 0.03) as illustrated in Figure 8a, with 96.7% similarity to its reference CP078863 (Figures 8b-c).
Figure 3. Identification and sequence analysis of vacA gene from H. pylori isolated biopsies. (a) Agarose gel electrophoresis of 2% agarose for the identification of vacA gene amplified at 259 bp from H. pylori isolated biopsies (b) vacA gene pairwise identity was 99.23% compared with the standard strain (CP079386) (c) The Blast Hit of vacA (259 bp) amplified area 662,662 → 662,920, relative to the original gene of the reference strain
Figure 4. Identification and sequence evaluation of the cagA gene derived from H. pylori isolated biopsies. (a) Agarose gel electrophoresis utilizing a 2% agarose medium to identify the cagA gene, which was amplified at 351 bp from H. pylori isolated biopsies. (b) The pairwise identity of the cagA gene was determined to be 96.3% in comparison to the reference strain (CP032037). (c) The Blast Hit of the cagA (351 bp) amplified region spans from 1,117,564 → 1,121,211, relative to the original gene of the reference strain
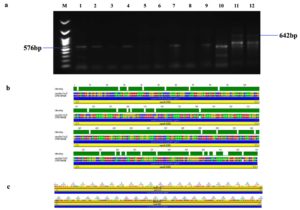
Figure 5. Identification and sequence analysis of vacA m1/m2 gene from H. pylori isolated biopsies. (a) Agarose gel electrophoresis of 1% agarose to identify vacA m1/m2 gene amplified at 576/642 bp from H. pylori isolated biopsies where M represents 100 bp ladder and lanes represent the positive samples (b) vacA m1/m2 gene pairwise identity was 92.11% compared with the standard strain (CP078468) (c) The Blast Hit of vacA m1/m2 (576/642 bp) amplified area 935,200 → 935,628, relative to the original gene of the reference strain
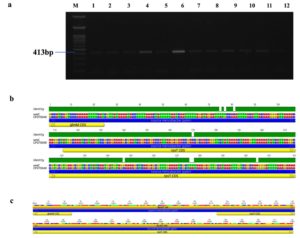
Figure 6. Identification and sequence analysis of ureC gene from H. pylori isolated biopsies. (a) Agarose gel electrophoresis of 1% agarose to identify ureC gene amplified at 413 bp from H. pylori isolated biopsies where M represents 100 bp ladder and lanes represent the positive samples. (b) ureC gene pairwise identity was 97.51% compared with the standard strain (CP079049) (c) The Blast Hit of ureC (413 bp) amplified area 74,601 → 74,921, relative to the original gene of the reference strain
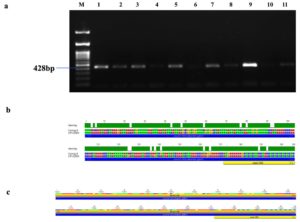
Figure 7. Identification and sequence analysis of oipA gene from H. pylori isolated biopsies. (a) Agarose gel electrophoresis of 1% agarose to identify oipA gene amplified at 428 bp from H. pylori isolated biopsies where M represents 100 bp ladder and lanes represent the positive samples. (b) oipA gene pairwise identity was 90.8% compared with the standard strain (C122953) (c) The Blast Hit of oipA (428 bp) amplified area 718,687 → 718,892, relative to the original gene of the reference strain
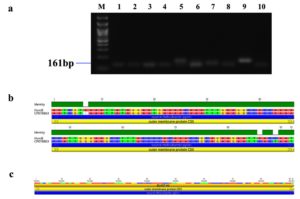
Figure 8. Identification and sequence analysis of homB gene from H. pylori isolated biopsies. (a) Agarose gel electrophoresis of 1% agarose to identify homB gene amplified at 161 bp from H. pylori isolated biopsies where M represents 100 bp ladder and lanes represent the positive samples. (b) Pairwise identity and nucleotide sequence for homB compared with the standard strain: CP078863 (NCBI) with 96.7% identity (c) Blast Hit of homB amplified region from 762,426 → 762,516, relative to the original gene of the reference strain
Clinical Impressions and their association with H. pylori genotypes
A higher frequency of H. pylori infection was detected for the virulence genes vacA s1 (71.89%), oipA (64.77%), vacA m1 (51.53%) and ureC (59.67%). The combination of vacA s1/m1 strain was 36.05% and vacA s1/m2 was 14.26%, as presented in Table 4. The vacA s1 was most common in adults between 18 to 35 years (24.9%; p = 0.048), while the cagA gene (39.61%; p = 0.02) and vacA m2 (15.38%; p = 0.036) was common among adults of 46 to 60 years (data not shown). It was inferred from our findings that the vacA s1 genotype has shown significant threat of acquiring gastritis (OR-1.68;95%;CI: 1.11-2.53; p = 0.013) and pangastritis (OR-4.79;95%;CI: 2.02-11.34; p = 0.0004) whereas our study population has reported that the vacA s1 genotype has reduced the risk for conditions such as gastric cancer (OR-0.13; 95%CI: 0.04-0.2; p = 0.0006), gastric ulcer (OR-0.10; 95%CI: 0.02-0.51; p = 0.005) and PUD (OR-0.27; 95% CI: 2.202-11.34; p = 0.01). In addition, the vacA s1/m1 combination strain has shown a higher incidence of gastritis (OR-2.34; 95%CI: 1.29-4.23; p = 0.004) and pangastritis (OR-2.61; 95%CI: 0.97-7.02; p = 0.05) and the vacA s1/m2 combination significantly correlated with gastritis (OR-2.03; 95%CI: 1.04-3.95; p = 0.03). The cagA strain showed a increased likelihood of GC (OR-3.45; 95%CI: 1.20-9.89; p = 0.02) and the oipA strain had a greater incidence of PUD (OR-7.87; 95%CI: 1.02-60.40; p = 0.04). The vacA m1, vacA m2, and oipA as well as vacA s1/m1 and vacA s1/m2 combinations showed no association with gastric cancer as shown in Table 4. Also, the vacA s1/m1 have shown lower risk for gastric ulcer (OR-0.06; 95%CI: 0.007-0.56; p = 0.01) whereas, homB gene had an increased incidence of gastritis (OR-1.62; 95%CI: 1.01-2.59; p = 0.04) and a markedly reduced risk for the development of duodenal ulcer (OR-0.17 95% CI: 0.04-0.72; p = 0.01).
Table (4):
Association of clinical impressions with H. pylori virulent genes
| Gene | Prevalence (%) | Clinical Impressions | ||||||
|---|---|---|---|---|---|---|---|---|
| Gastritis | Duodenal Ulcer | Gastric cancer | Gastric ulcer | Pangastritis | PUD | Nodular Gastritis | ||
| OR (95% CI) | ||||||||
| vacA s1 | 71.89 | 1.68 (1.11-2.53) 0.013 | 0.52 (0.25-1.03) 0.063 | 0.13 (0.04-0.42) 0.0006 | 0.10 (0.02-0.51) 0.005 | 4.79 (2.02-11.34) 0.0004 | 0.27 (0.05-1.42) 0.01 | 1.11 (0.12-11.38) 0.9 |
| cagA | 31.16 | 0.71 (0.48-1.07) 0.105 | 1.11 (0.54-2.28) 0.77 | 3.45 (1.20-9.89) 0.02 | 2.82 (0.074-10.65) 0.126 | 0.89 (0.52-1.51) 0.668 | 1.97 (0.70-5.55) 0.195 | 2.22 (0.31-15.94) 0.426 |
| vacA m1 | 51.53 | 1.13 (0.77-1.65) 0.509 | 1.34 (0.67-2.67) 0.105 | 0.22 (0.06-0.81) 0.028 | 0.46 (0.11-1.87) 0.281 | 1.01 (0.62-1.64) 0.942 | 1.42 (0.49-4.06) 0.105 | 0.31 (0.03-3.00) 0.313 |
| vacA m2 | 21.18 | 1.15 (0.71-1.84) 0.56 | 1.26 (0.57-1.26) 0.56 | 0.91 (0.25-3.31) 0.89 | 3.05 (0.80-11.59) 0.1 | 0.79 (0.42-1.48) 0.46 | 0.56 (0.12-2.54) 0.1 | – |
| ureC | 59.67 | 1.01 (0.68-1.48) 0.955 | 1.38 (0.67-2.83) 0.376 | 1.89 (0.59-6.02) 0.28 | 1.35 (0.33-5.49) 0.667 | 0.72 (0.44-1.18) 0.199 | 1.01 (0.35-2.89) 0.105 | 0.67 (0.09-4.82) |
| oipA | 64.77 | 1.05 (0.71-1.56) 0.78 | 1.16 (0.56-2.37) 0.68 | 0.18 (0.05-0.59) 0.004 | – | 0.55 (0.34-0.91) 0.01 | 7.87 (1.02- 60.40) 0.04 | – |
| homB | 24.03 | 1.62 (1.01-2.59) 0.04 | 0.17 (0.04-0.72) 0.01 | 0.78 (0.21-2.82) 0.711 | – | 1.37 (0.80-2.34) 0.24 | 0.21 (0.02-1.68) 0.14 | – |
| vacA s1/m1 | 36.05 | 2.34 (1.29-4.23) 0.004 | 0.75 (0.25-2.26) 0.61 | 0.08 (0.02-0.29) 0.0002 | 0.06 (0.007-0.56) 0.01 | 2.61 (0.97-7.02) 0.05 | 0.02 (0.003-0.18) 0.0004 | 0.34 (0.02-5.62) 0.45 |
| vacA s2/m2 | 14.26 | 2.03 (1.04-3.95) 0.03 | 0.57 (0.17-1.89) 0.51 | 0.24 (0.05-1.15) 0.07 | – | 1.74 (0.72-4.22) 0.21 | – | – |
OR odds ratio was carried out to establish the association between clinical conditions with H. pylori virulent genes. % denotes the percentage of each virulent gene present in the patients. 95% CI: 95% Confidence interval. Values highlighted represent the statistical significance at the p <0.05 level.
H. pylori, a challenging bacterium to cultivate and identify due to its meticulous nature, which requires significant time and effort. Therefore, molecular techniques that offer high sensitivity and specificity for detecting these bacteria are becoming increasingly popular for studying their prevalence in various experimental samples.27 The diverse genetic makeup of virulence factors among H. pylori strains worldwide contributes to a broad spectrum of gastrointestinal diseases. Increasing evidence suggests that H. pylori genetic diversity has significant clinical implications.28 The present study provides a comprehensive molecular and histopathological evaluation of H. pylori infection in gastric biopsy tissues by employing RUT, Giemsa staining and PCR amplification by 16S rRNA gene. The identification of H. pylori in 98.2% of the samples by PCR underscores a greater sensitivity and specificity of molecular strategies compared to conventional diagnostic methods. Our findings were consistent with previous studies conducted by Gantuya et al., who reported 94.3% detection rate of H. pylori using 16S rRNA specific PCR and Han et al. observed 92.2% positivity rate.29,30 Similarly, Elnosh et al. reported a 78.6% detection rate, emphasizing that PCR targeting the 16S rRNA gene was one of the most reliable results for molecular detection of H. pylori.31
H. pylori, the most genetically diverse bacterial pathogen, exhibiting variable virulence genotypes across populations from different geographic origins. Molecular typing methods have significantly enhanced our insights into the epidemiology of H. pylori infection.32 In our study, various virulence genes of H. pylori were analyzed, for the presence of virulence-associated genes, particularly vacA (s1, m1/m2), cagA, oipA, ureC and homB.
The vacA gene, an important virulence factor, displays a mosaic structure characterized by two groups of allelic variations: the signal sequence (s1, s2) and the mid-region (m1, m2).33 Extensive investigations have been conducted on allelic variation as a virulence determinant in H. pylori strains. All H. pylori strains harbour the vacA gene; however, the difference in the signal and mid-regions alter cytotoxicity. The variation in vacA genotypes (allelic variations) was notably distinct across different countries, with prior studies highlighting significant geographic disparities in these virulence factors.23 In our investigation, the vacA s1 genotype was predominant in 71.89% of cases, which was consistent with previous reports highlighting its strong association with gastroduodenal diseases. Notably, vacA m1 strains, which have been implicated in increased cytotoxic activity and enhanced gastric epithelial cell vacuolation, were reported in 51.53% of cases, while the vacA s1/m1 was reported in 36.05% and the vacA s1/m2 was reported in 14.26% of our cases. Hence, these findings have revealed the higher incidence of vacA gene among H. pylori-infected patients which aligns with studies conducted in various regions of India such as Hyderabad, Varanasi, Coimbatore and Chennai.34-36 Similarly, our study correlated with studies reported for H. pylori virulence toxins vacA s1 exhibiting high prevalence rates in Asia, South Africa, Northern Europe, South America, and Southern Europe37,38 and studies conducted in Brazil and the Middle East indicate that vacA s1 or m1 genotypes are linked to the advancement of stomach cancer.39 Also, Mukhopadhyay et al. observed notable prevalence of the vacA s1m1 allele in Kolkata.40 In addition, the findings of our study aligns with the research conducted on patients in the Khuzestan province of Iran, which reported higher vacA s1/m1 genotypes.33 Nonetheless, a study by Udayakumar et al. has indicated that the vacA m2 allele was more commonly reported in Chennai.34 Moreover, our findings have discovered notable correlations between particular H. pylori genotypes and a range of gastrointestinal disorders where our analysis revealed that the association of vacA s1 with specific gastric conditions was predominantly noted in patients diagnosed with gastritis (OR = 1.68; 95% CI 1.11-2.53; p = 0.013) and pangastritis (OR = 4.79; 95% CI 2.02-11.34; p = 0.0004), indicating increased odds of these inflammatory conditions in individuals harboring s1-positive strains. Similarly, vacA s1/m1 genotypes demonstrated increased odds of gastritis (OR – 2.34; 95%CI: 1.29-4.23; p = 0.004) and pangastritis (OR – 2.61; 95%CI: 0.97-.02; p = 0.05), indicating its contribution to mucosal inflammation. These results align with previous studies involving Iranian and South Korean populations.41,42 Also, the vacA s1/m2 genotype was significantly associated with increased odds of gastritis (OR – 2.03; 95% CI: 1.04-3.95; p = 0.03) and inversely linked with other conditions and they align with previous studies conducted.23,34,40 Interestingly, although vacA genotypes are globally associated with more severe disease outcomes such as duodenal ulcer, gastric cancer and gastric ulcer our study exhibited a lower risk of these conditions among vacA-positive individuals. This inverse association was likely influenced by very small sample sizes among these gastric conditions, leading to imprecise odds ratio estimates and wide confidence intervals. Therefore, vacA being an important virulence marker for the gastrointestinal conditions in our South Indian population, these findings should be interpreted with caution. Furthermore, the results highlight how the pathogenic potential of globally significant virulence factors can manifest differently across specific regional populations.
The cagA gene was identified as a notable virulence factor, potentially elevating the risk of drug resistance and carcinoma in affected individuals. The correlation between cag PAI and infection status in East Asia was approximately 60%, whereas in India, it is around 90%. However, our study found that the cagA genotype, an indicator for cag PAI presence, was 31.36% indicating a moderate prevalence within the studied population and was significantly associated with an elevated risk of GC (OR-3.45; 95%CI: 1.20-9.89, p ≤ 0.02), reinforcing its role as a critical virulence determinant in H. pylori-induced carcinogenesis. These findings were similar to a study on the comparative analysis of H. pylori isolates from North and South India, which demonstrated significant variations in the integrity of the cagA gene and cagPAI between the two regions. Approximately 34% of the isolates obtained from Varanasi and 46% of the isolates from Hyderabad exhibited the complete cagPAI. In comparison, 65.2% of the isolates from Varanasi exhibited a partially deleted cagPAI, while 53.5% of the isolates from Hyderabad showed the same characteristic features. The frequency of this gene exhibits considerable variation worldwide, with reported values between 60% and 80%.43,44 However, numerous studies have reported that the cagA gene has been linked to gastritis, PUD and duodenal ulcer; however, our findings have discovered that there was a lower correlation with any of these gastric conditions.45,46 The findings reinforce the critical role of cagA as a virulence determinant in H. pylori-induced carcinogenesis, even when its overall prevalence is lower in our population. Although, cagA has been widely associated with gastritis, duodenal ulcer in many populations, our study did not observe such correlations. This discrepancy likely reflects the regional variations in cagA gene that includes EPIYA motifs or partial deletion within the pathogenicity island which could diminish its functional activity. The characterization of the predominant circulating strains and their disease associations was indispensable for developing region-specific clinical guidelines and contributes to the global understanding of the H. pylori virulence markers. Hence, the establishment of this genotypic baseline provides the essential scientific framework for future mechanistic studies emphasizing South Indian strains.47-49
The identification of ureC, a prerequisite for bacterial survival in acidic environments, was observed in 59.6% of cases, underscoring the adaptability of H. pylori within the gastric niche. Our study has reported a higher prevalence of the ureC gene in comparison with a study carried out in Gujarat, India, where they reported 18.18% of H. pylori-positive samples with ureC gene and there was no correlation with the gastric conditions.50 However, a study conducted in Taiwan showed that the MboI-defined genotype 3 ureC gene exhibits a closer association with gastric cancer and gastric ulcer. Thus, the genetic variations in ureC may impact the pathogenesis of gastric diseases.14
oipA and homB are well-known outer inflammatory proteins (OMPs) of H. pylori that play a crucial role for immune response stimulation, cell attachment, and initiation of infection. oipA, an important adhesion associated with bacterial colonization and inflammation, was identified in 64.97% of cases and showed a correlation with PUD (OR – 7.87; 95%CI: 1.02-60.40; p = 0.04) in our study population however, this extremely wide confidence interval indicates low precision due to decreased sample size of PUD cases, as that of the study reported by Rezaei et al., with the prevalence of 56% oipA gene. In addition, the study investigated by El-Sayed et al. in the Egyptian population, where the oipA virulence gene showed a higher association with gastroduodenitis.9,51 However, the oipA gene reported by Singh et al. and Esteghamati et al. have found no significant association with distinct gastrointestinal diseases such as DU, GERD, NERD and gastritis & PUD respectively. While oipA was identified at a high frequency and demonstrated a significant association with PUD in our population, this finding must be approached with caution due to the limited number of PUD cases, which led to a very wide confidence interval and restricted statistical precision. The variability in the functional ‘on/off’ status of oipA, along with regional strain diversity and host-specific inflammatory responses, may contribute to the inconsistencies observed across studies. Despite these limitations, oipA was recognized as a significant mediator of gastric inflammation, and our findings suggest that it may serve as a virulence biomarker in patients with PUD. Therefore, comprehensive studies with broader spectrum of ulcer cases are necessary to elucidate their accurate clinical significance among the South Indian population and to facilitate the formulation of targeted antimicrobial strategies.52,53
Our findings are concordant with previous studies by Oleastro et al. in the French population and Bajwa et al. in the Pakistani population, which showed a significant association with PUD and gastritis for the prevalence of homB, where we reported 24.03% and others reported 58% and 38.8%, respectively. Consistent with earlier studies, we observed that homB-positive strains were more common in patients with gastritis (OR – 1.62; 95%CI: 1.01-2.59; p = 0.04). 21,54 However, in our study, the homB did not show any significant correlation with GC, unlike the study reported by Talebi et al., in Iranian population. This discrepancy could indicate regional variations in circulating H. pylori lineages, differences in the distribution of homA/homB paralogues, or host immune responses that influence the expression the gene. The limited number of gastric cancer cases in our cohort likely restricted the statistical power to identify associations with less prevalent virulence markers. Although homB appears to contribute for inflammatory diseases in our population, its involvement in malignant outcomes necessitates additional research through larger, multi-centre studies.55
Therefore, in our study we found that the presence of specific virulence genes significantly influences the development of various gastrointestinal diseases and PCR-based detection in our study revealed diverse distributions of vacA, cagA, oipA, homB and ureC genotypes across various gastric conditions, highlighting the importance of genotype-specific risk assessment to guide clinical management and treatment strategies. Consequently, the genotypic diversity identified among the H. pylori isolates suggests that various strains with differing pathogenic potential may exist within a population, leading to disease heterogeneity.
Limitations
Our study was conducted in a single medical facility; therefore, the potential for selection bias cannot be rule out. As this study was confined to the South Indian Tamil population, the ability to extrapolate its findings to a wider range of global cohorts was limited. Although several significant associations were observed between the virulence genotypes and clinical outcomes, the unequal distribution of cases across clinical outcomes may have contributed to highly variable odds ratio estimates and wide confidence intervals, warranting cautious interpretation of genotype-disease associations. Furthermore, the cross-sectional design restricts the ability to infer causality between the presence of virulence genes and disease progression. The highly sensitive PCR-based molecular method detected only the presence of virulence genes and did not evaluate their expression levels, functional activity, host-pathogen interaction or genotype-phenotype patterns, which could provide a more precise indication of pathogenic potential. The inability to characterize mixed infections or allelic diversity- common in H. pylori populations also represents an additional limitation. These constraints provide essential context for interpretation and highlight the need for larger, multicentric, and mechanistic studies to enable the inclusion of appropriate control groups in near future.
H. pylori strains demonstrated significant genotypic diversity, with vacA s1 identified as the most common virulence genotype in our population. Significant associations were identified between vacA s1 and vacA s1/m1 genotypes with gastritis and pangastritis, while cagA showed a strong correlation with gastric cancer. The prevalence of virulence factor genes was reported in the increasing order of vacA s1 > oipA > ureC > vacA m1 > vacA s1/m1 > cagA > vacA m2 > vacA s1/m2, from isolates of patients infected with H. pylori infection. These findings suggest that vacAs1 can may serve as one of the potent virulent markers for gastrointestinal disease manifestation. Also, our findings highlight the genetic diversity among H. pylori strains and their effects on gastric pathology. However, given the cross-sectional design and limited subgroups sizes, the association should be interpreted cautiously. Further multicentre and longitudinal studies, including gene expression analyses are needed to emphasizes the pathogenic significance of genotype and to enlighten clinical management and treatment strategies for H. pylori-associated gastrointestinal diseases.
ACKNOWLEDGMENTS
The authors would like to thank the Department of Genetic Engineering, School of Bioengineering, SRM Institute of Science and Technology, Kattankulathur for assistance with research facilities.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
MJ conceptualized and designed the study. NAR recruited the cases for the study and performed endoscopies. SKSR performed sample collection. SKSR performed the experiments. SKSR and MJ performed data analysis. MJ and SKSR wrote the manuscript. MJ reviewed and edited the manuscript. All authors read and approved the final manuscript for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This study was approved by the Institutional Ethical Committee, SRM Medical College and Research Center, Kattankulathur, Chennai, India, dated 25/03/2021 with EC Reg. No. 2394/IEC/2021.
INFORMED CONSENT
Written informed consent was obtained from the participants before enrolling in the study.
- Kao CY, Sheu BS, Wu JJ. Helicobacter pylori infection: An overview of bacterial virulence factors and pathogenesis. Biomed J. 2016;39(1):14-23.
Crossref - Burucoa C, Axon A. Epidemiology of Helicobacter pylori infection. Helicobacter. 2017;22(S1):e12403.
Crossref - Kharel S, Bist A, Shrestha S, Homagain S. Helicobacter pylori healthy South Asians. JGH Open. 2020;4(6):1037-1046.
Crossref - Puzhakkal S, Mittal P, Thiruchelvam K. Prevalence of Helicobacter pylori Infection in India: A Systematic Re-view and Meta-Analysis. Evidence Synthesis in Healthcare Connect. 2025;1(1):1-10.
Crossref - Nishida T, Tsujii Y, Okamoto A, et al. A Triple-Drug Blister-Packaged Drug with Vonoprazan Improves First-Line Eradication of Helicobacter pylori in Elderly Patients: A Retrospective Propensity Score-Matched Cohort Study. Digestion. 2020;101(5):608-614.
Crossref - Sharndama HC, Mba IE. Helicobacter pylori: an up-to-date overview on the virulence and pathogenesis mechanisms. Braz J Microbiol. 2022;53(1):33-50.
Crossref - Maeda M, Moro H, Ushijima T. Mechanisms for the induction of gastric cancer by Helicobacter pylori infection: aberrant DNA methylation pathway. Gastric Cancer. 2017;20(1):8-15.
Crossref - Souod N, Kargar M, Doosti A, Ranjbar R, Sarshar M. Genetic Analysis of cagA and vacA Genes in Helicobacter Pylori Isolates and Their Relationship with Gastroduodenal Diseases in the West of Iran. Iran Red Crescent Med J. 2013;15(5):371-375.
Crossref - Rezaei F, Alebouyeh M, Mirbagheri SZ, Ebrahimi A, Foroushani AR, Bakhtiari R. Transcriptional analysis of Helicobacter pylori cytotoxic-associated gene-pathogenicity island in response to different pH levels and proton pump inhibitor exposure. Indian J Gastroenterol. 2023;42(5):686-693.
Crossref - Foegeding NJ, Caston RR, McClain MS, Ohi MD, Cover TL. An Overview of Helicobacter pylori VacA Toxin Biology. Toxins. 2016;8(6):173.
Crossref - Matsuo Y, Kido Y, Yamaoka Y. Helicobacter pylori Outer Membrane Protein-Related Pathogenesis. Toxins. 2017;9(3):101.
Crossref - Liu J, He C, Chen M, Wang Z, Xing C, Yuan Y. Association of presence/absence and on/off patterns of Helicobacter pylori oipA gene with peptic ulcer disease and gastric cancer risks: a meta-analysis. BMC Infect Dis. 2013;13(1):1-10.
Crossref - De Reuse H, Labigne A, Mengin-Lecreulx D. The Helicobacter pylori ureC gene codes for a phosphoglucosamine mutase. J Bacteriol. 1997;179(11):3488-3493.
Crossref - Chen TH, Cheng HT, Yeh CT. Epidemiology changes in peptic ulcer diseases 18 years apart explored from the genetic aspects of Helicobacter pylori. Translational Research. 2021;232:115-120.
Crossref - Alm RA, Bina J, Andrews BM, Doig P, Hancock REW, Trust TJ. Comparative Genomics of Helicobacter pylori: Analysis of the Outer Membrane Protein Families. Infect Immun. 2000;68(7):4155-4168
- World Cancer Research Fund International. Stomach cancer statistics. WCRF. https://www.wcrf.org/preventing-cancer/cancer-statistics/stomach-cancer-statistics/. Accessed May 15, 2025.
- Chen TS, Li FY, Chang FY, Lee SD. Immunoglobulin G antibody against Helicobacter pylori: clinical implications of levels found in serum. Clin Diagn Lab Immunol. 2002;9(5):1044-1048.
Crossref - Ghosh P, Sarkar A, Ganguly M, et al. Helicobacter pylori strains harboring babA2 from Indian sub population are associated with increased virulence in ex vivo study. Gut Pathog. 2016;8:1.
Crossref - Eslaminejad A, Marashian SM, Aboutorabi M, Sadr M, Agah S. Determination of optimal time for reading of rapid urease test diagnosis of Helicobacter pylori. Gastroenterol Hepatol Bed Bench. 2020;13(3):232-237.
- Kim J, Lee S, Lee SP, et al. The histologic detection of Helicobacter pylori in seropositive subjects is affected by pathology and secretory ability of the stomach. Helicobacter. 2018;23(3):e12480.
Crossref - Oleastro M, Cordeiro R, Ferrand J, et al. Evaluation of the Clinical Significance of homB a Novel Candidate Marker of Helicobacter pylori Strains Associated with Peptic Ulcer Disease. J Infect Dis. 2008;198(9):1379-1387.
Crossref - Idris AB, Hassan HG, Ali MAS, et al. Molecular Phylogenetic Analysis of 16S rRNA Sequences Identified Two Lineages of Helicobacter pylori Strains Detected from Different Regions in Sudan Suggestive of Differential Evolution. Int J Microbiol. 2020;2020:8825718.
Crossref - Atherton JC, Cao P, Peek RM, Tummuru MKR, Blaser MJ, Cover TL. Mosaicism in Vacuolating Cytotoxin Alleles of Helicobacter pylori: Association of specific vaca types with cytotoxin production and peptic ulceration. J Biol Chem. 1995;270(30):17771-17777.
Crossref - Chattopadhyay S, Patra R, Ramamurthy T, et al. Multiplex PCR Assay for Rapid Detection and Genotyping of Helicobacter pylori Directly from Biopsy Specimens. J Clin Microbiol. 2004;42(6):2821-2824.
Crossref - Javanbakhat P, Peeridogaheh H, Nemati R, et al. Exploring virulence factors of Helicobacter pylori isolated from gastric biopsy. Mol Biol Rep. 2024;51(1):192.
Crossref - Yamaoka Y, Kwon DH, Graham DY. A Mr 34,000 proinflammatory outer membrane protein (oipA) of Helicobacter pylori. Proc Natl Acad Sci U S A. 2000;97(13):7533-7538.
Crossref - Godbole G, Megraud F, Bessede E. Review: Diagnosis of Helicobacter pylori infection. Helicobacter. 2020;25(Suppl 1):e12735.
Crossref - Shiota S, Suzuki R, Yamaoka Y. The significance of virulence factors in Helicobacter pylori. J Digest Dis. 2013;14(7):341-349.
Crossref - Han HS, Lee KY, Lim SD, Kim WS, Hwang TS. Molecular identification of Helicobacter DNA in human gastric adenocarcinoma tissues using Helicobacter species-specific 16S rRNA PCR amplification and pyrosequencing analysis. Oncol Lett. 2010;1(3):555-558.
Crossref - Gantuya B, Serag HBE, Saruuljavkhlan B, et al. Advantage of 16S rRNA Amplicon Sequencing in Helicobacter pylori Diagnosis. Helicobacter. 2021;26(3):e12790.
Crossref - Elnosh M, Altayb H, Hamedelnil Y, et al. Comparison of invasive histological and molecular methods in the diagnosis of Helicobacter pylori from gastric biopsies of Sudanese patients: a cross-sectional study. F1000Res. 2022;11:113.
Crossref - Yamaoka Y. Mechanisms of disease: Helicobacter pylori virulence factors. Nat Rev Gastroenterol Hepatol. 2010;7(11):629-641.
Crossref - Sheikh AF, Yadyad MJ, Goodarzi H, et al. CagA and vacA allelic combination of Helicobacter pylori in gastroduodenal disorders. Microb Pathogen. 2018;122:144-150.
Crossref - Udhayakumar G, Senthilkumar C, Jayanthi V, Devaraj N, Devaraj H. Helicobacter pylori detection and genotyping in gastric biopsy specimens from Chennai patients (India). Can J Microbiol. 2009;55(2):126-132.
Crossref - Kumar S, Kumar A, Dixit VK. Genetic diversity in strains of Helicobacter pylori from India and their relatedness to strains from other parts of the world. Infection, Genetics and Evolution. 2011;11(1):242-247.
Crossref - Jeyamani L, Jayarajan J, Leelakrishnan V, Swaminathan M. CagA and VacA genes of Helicobacter pylori and their clinical relevance. Indian J Pathol Microbiol. 2018;61(1):66-69.
Crossref - van Doorn LJ, Figueiredo C, Rossau R, et al. Typing of Helicobacter pylori vacA gene and detection of cagA gene by PCR and reverse hybridization. J Clin Microbiol. 1998;36(5):1271-1276.
Crossref - Yamaoka Y, Kodama T, Gutierrez O, Kim JG, Kashima K, Graham DY. Relationship between Helicobacter pylori iceA, cagA, and vacA Status and Clinical Outcome: Studies in Four Different Countries. J Clin Microbiol. 1999;37(7):2274-2279. Accessed September 7, 2023. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC85136/
- Sarma A, Saikia L, Gogoi M, Yadav K. Molecular Characterisation of Virulent Gene vacA in Helicobacter pylori Clinical Isolates from Patients with Gastroduodenal Diseases in Assam, India. Indian J Med Microbiol. 2018;36(2):178-185.
Crossref - Mukhopadhyay AK, Kersulyte D, Jeong JY, et al. Distinctiveness of Genotypes of Helicobacter pylori in Calcutta, India. J Bacteriol. 2000;182(11):3219-3227. Accessed April 14, 2024.
Crossref - Kim JY, Kim N, Nam RH, et al. Association of polymorphisms in virulence factor of Helicobacter pylori and gastroduodenal diseases in South Korea. J Gastroenterol Hepatol. 2014;29(5):984-991.
Crossref - Matos JI, de Sousa HAC, Marcos-Pinto R, Dinis-Ribeiro M. Helicobacter pylori CagA and VacA genotypes and gastric phenotype: a meta-analysis. Eur J Gastroenterol Hepatol. 2013;25(12):1431.
Crossref - Kumar S, Kumar A, Dixit VK. Diversity in the cag pathogenicity island of Helicobacter pylori isolates in populations from North and South India. J Med Microbiol. 2010;59(1):32-40.
Crossref - Shetty V, Lingadakai R, Pai GC, Ballal M. Profile of Helicobacter pylori cagA & vacA genotypes and its association with the spectrum of gastroduodenal disease. Indian J Med Microbiol. 2021;39(4):495-499.
Crossref - Khadir ME, Boukhris SA, Benajah DA, et al. VacA and CagA Status as Biomarker of Two Opposite End Outcomes of Helicobacter pylori Infection (Gastric Cancer and Duodenal Ulcer) in a Moroccan Population. PLOS ONE. 2017;12(1):e0170616.
Crossref - Bustos-Fraga S, Salinas-Pinta M, Vicuna-Almeida Y, de Oliveira RB, Baldeon-Rojas L. Prevalence of Helicobacter pylori genotypes: cagA, vacA (m1), vacA (s1), babA2, dupA, iceA1, oipA and their association with gastrointestinal diseases. A cross-sectional study in Quito-Ecuador. BMC Gastroenterol. 2023;23:197.
Crossref - Malfertheiner P, Megraud F, O’Morain CA, et al. Management of Helicobacter pylori infection-the Maastricht V/Florence Consensus Report. Gut. 2017;66(1):6-30.
Crossref - Hooi JKY, Lai WY, Ng WK, et al. Global Prevalence of Helicobacter pylori Infection: Systematic Review and Meta-Analysis. Gastroenterology. 2017;153(2):420-429.
Crossref - Cover TL, Lacy DB, Ohi MD. The Helicobacter pylori Cag Type IV Secretion System. Trends Microbiol. 2020;28(8):682-695.
Crossref - Pandya HB, Agravat HH, Patel JS. Prevalence of Specific Helicobacter Pylori cagA, vacA, iceA, ureC Genotypes and its Clinical Relevance in the Patients with Acid-Peptic Diseases. J Clin Diagn Res. 2017;11(8):DC23-DC26.
Crossref - El-Sayed MS, Musa N, Eltabbakh M, et al. Detection of Helicobacter pylori oipA and dupA genes among dyspeptic patients with chronic gastritis. Alexandria Journal of Medicine. 2020;56(1):105-110.
Crossref - Singh S, Sharma P, Mahant S, Das K, Som A, Das R. Analysis of Functional Status of Genetically Diverse OipA Gene in Indian Patients with Distinct Gastrointestinal Disease. Curr Microbiol. 2022;80(1):35.
Crossref - Esteghamati A, Sayyahfar S, Khanaliha K, et al. Prevalence and Clinical Relevance of cagA and oipA Genotypes of Helicobacter pylori in Children and Adults with Gastrointestinal Diseases in Tehran, Iran. Med J Islam Repub Iran. 2023;37(1):169-173.
Crossref - Bajwa MA, Idrees M, Maheshwari PK. Association of dupA, iceA, homB Genes of Helicobacter pylori with Gastritis, Peptic Ulcer Disease and Gastric Cancer. J Pharma Res Int. 2020;32(25):1-6.
Crossref - Abadi ATB, Rafiei A, Ajami A, et al. Helicobacter pylori homB, but not cagA, is associated with gastric cancer in Iran. J Clin Microbiol. 2011;49(9):3191-3197.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.